Deposition Date
2004-06-03
Release Date
2005-03-02
Last Version Date
2023-12-13
Entry Detail
PDB ID:
1W19
Keywords:
Title:
Lumazine Synthase from Mycobacterium tuberculosis bound to 3-(1,3,7- trihydro-9-D-ribityl-2,6,8-purinetrione-7-yl)propane 1-phosphate
Biological Source:
Source Organism(s):
MYCOBACTERIUM TUBERCULOSIS (Taxon ID: 1773)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.00 Å
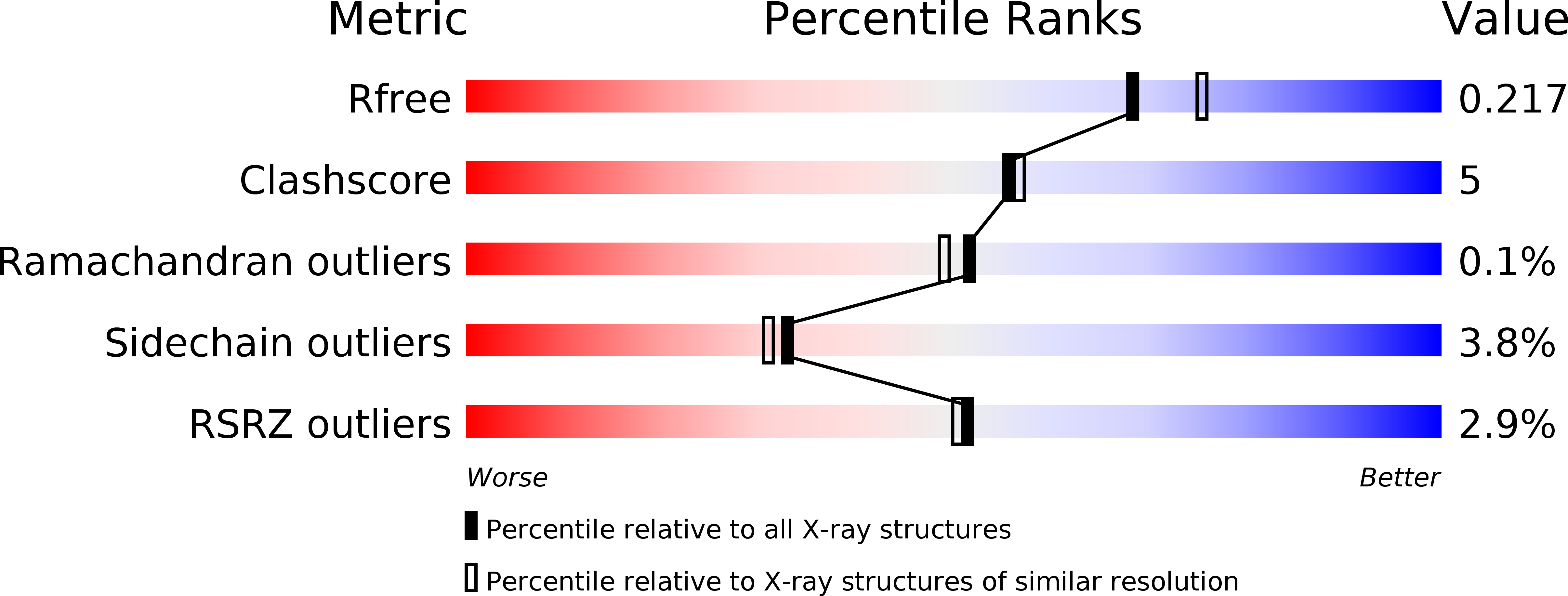
R-Value Free:
0.19
R-Value Work:
0.14
R-Value Observed:
0.15
Space Group:
C 1 2 1


