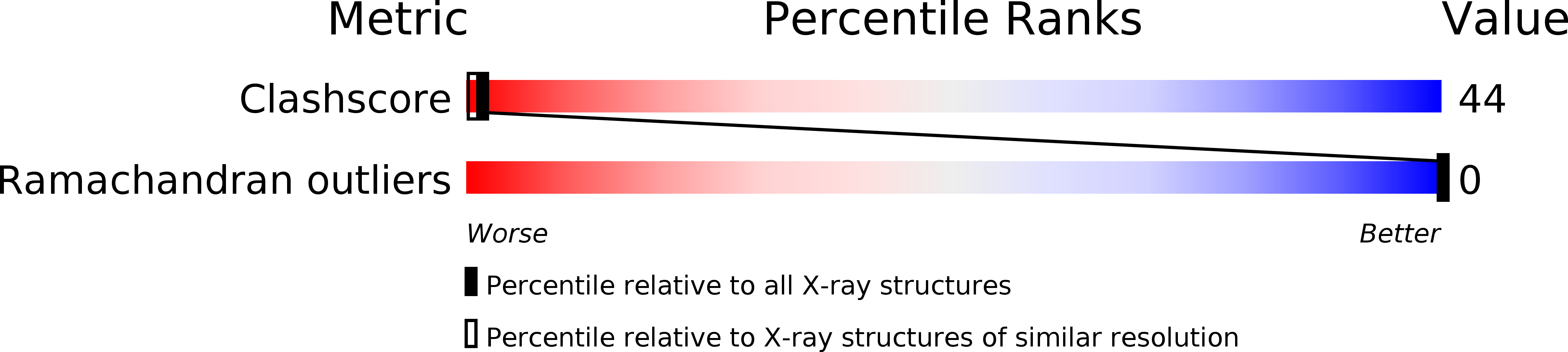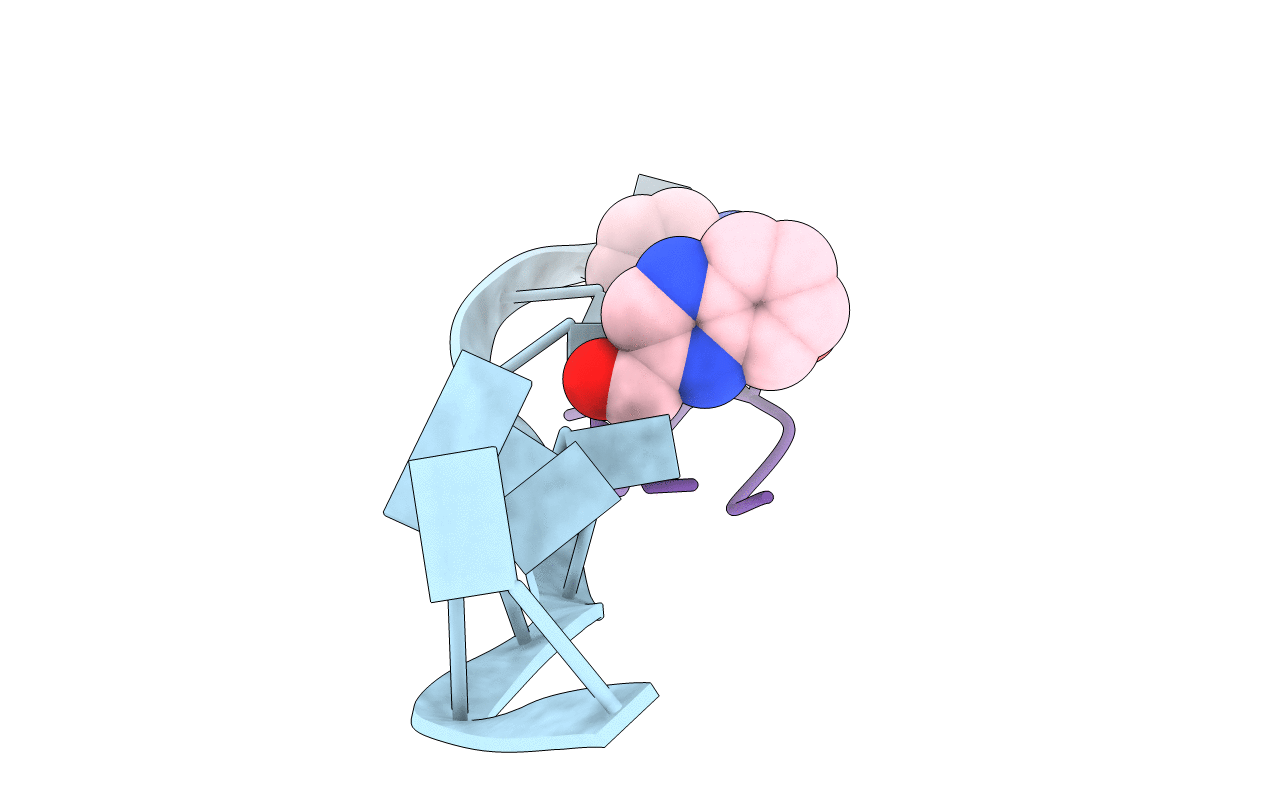
Deposition Date
1986-10-21
Release Date
2006-06-27
Last Version Date
2023-12-27
Entry Detail
PDB ID:
1VS2
Keywords:
Title:
Interactions of quinoxaline antibiotic and DNA: the molecular structure of a TRIOSTIN A-D(GCGTACGC) complex
Biological Source:
Source Organism(s):
STREPTOMYCINEAE (Taxon ID: 1931)
Method Details:
Experimental Method:
Resolution:
2.00 Å
R-Value Observed:
0.20
Space Group:
P 63 2 2