Abstact
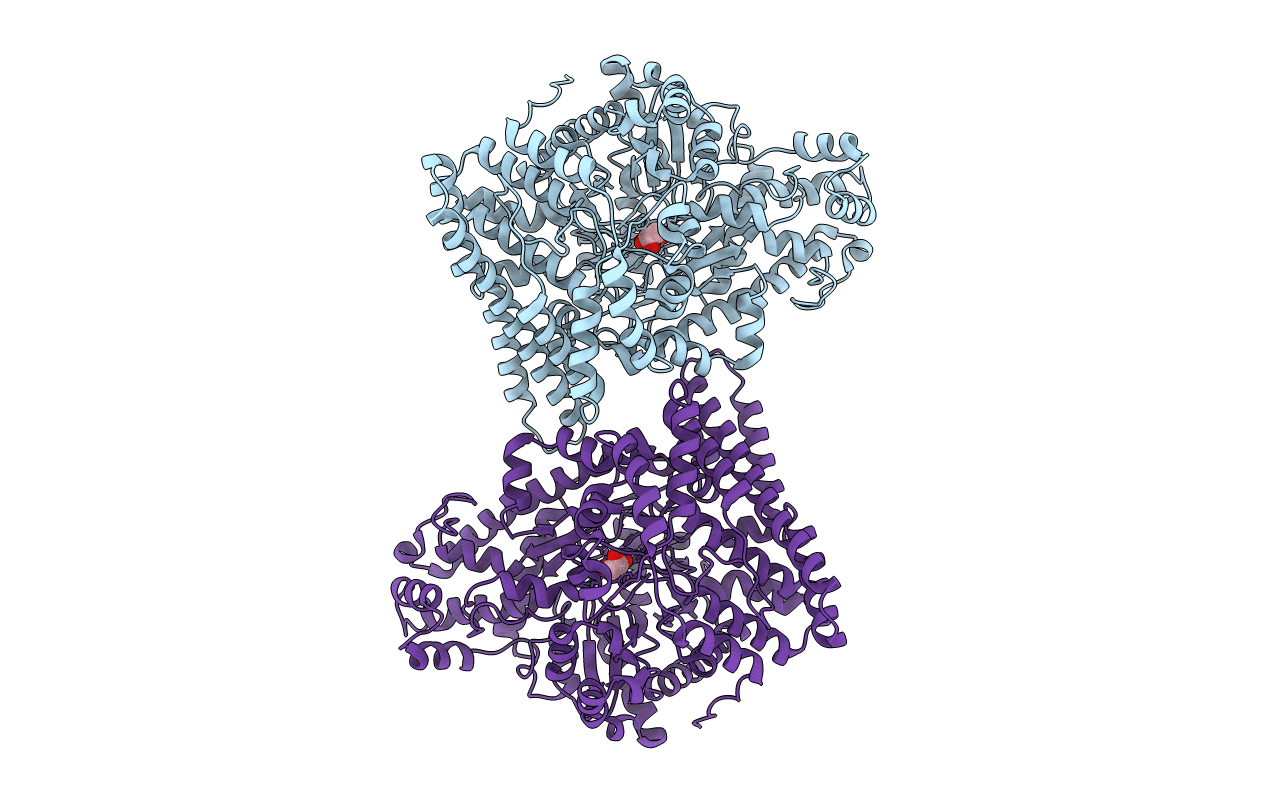
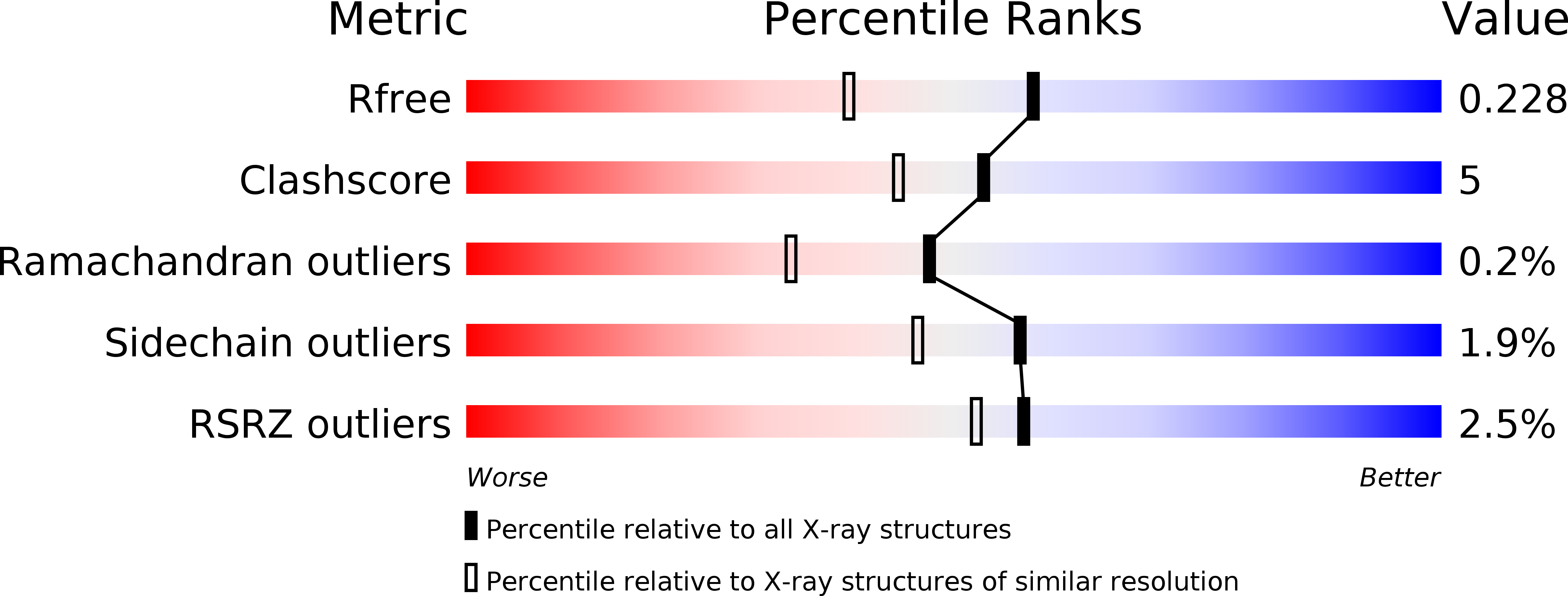
The molecular characterization of a B12-independent glycerol dehydratase from Clostridium butyricum has recently been reported [Raynaud, C., et al. (2003) Proc. Natl. Acad. Sci. U.S.A. 100, 5010-5015]. In this work, we have further characterized this system by biochemical and crystallographic methods. Both the glycerol dehydratase (GD) and the GD-activating enzyme (GD-AE) could be purified to homogeneity under aerobic conditions. In this form, both the GD and GD-AE were inactive. A reconstitution procedure, similar to what has been reported for pyruvate formate lyase activating enzyme (PFL-AE), was employed to reconstitute the activity of the GD-AE. Subsequently, the reconstituted GD-AE could be used to reactivate the GD under strictly anaerobic conditions. We also report here the crystal structure of the inactive GD in the native (2.5 A resolution, Rcryst = 17%, Rfree = 20%), glycerol-bound (1.8 A resolution, Rcryst = 21%, Rfree = 24%), and 1,2-propanediol-bound (2.4 A resolution, Rcryst = 20%, Rfree = 24%) forms. The overall fold of the GD monomer was similar to what has been observed for pyruvate formate lyase (PFL) and anaerobic ribonucleotide reductase (ARNR), consisting of a 10-stranded beta/alpha barrel motif. Clear density was observed for both substrates, and a mechanism for the dehydration reaction is presented. This mechanism clearly supports a concerted pathway for migration of the OH group through a cyclic transition state that is stabilized by partial protonation of the migrating OH group. Finally, despite poor alignment (rmsd approximately 6.8 A) of the 10 core strands that comprise the barrel structure of the GD and PFL, the C-terminal domains of both proteins align well (rmsd approximately 0.7 A) and have structural properties consistent with this being the docking site for the activating enzyme. A single point mutation within this domain, at a strictly conserved arginine residue (R782K) in the GD, resulted in formation of a tight protein-protein complex between the GD and the GD-AE in vivo, thereby supporting this hypothesis.