Deposition Date
2003-07-14
Release Date
2003-09-02
Last Version Date
2024-11-06
Entry Detail
PDB ID:
1PZY
Keywords:
Title:
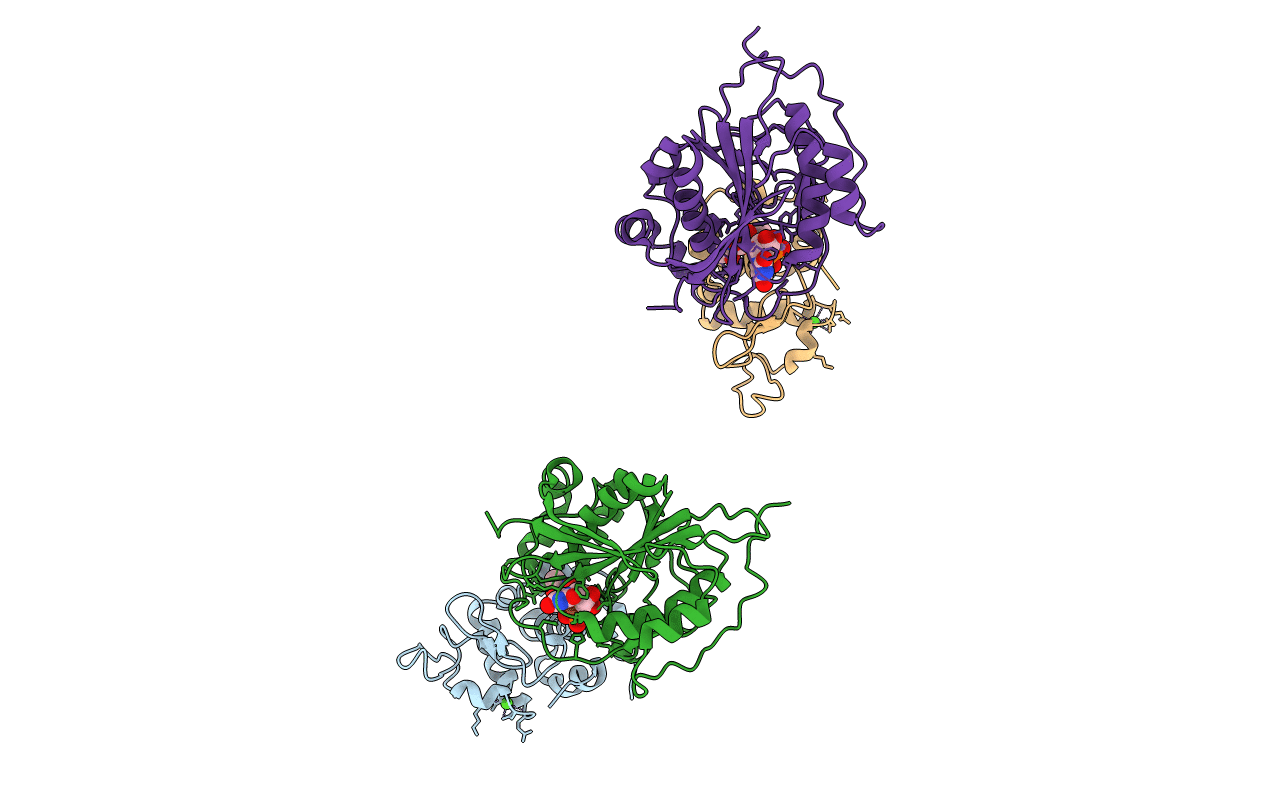
W314A-BETA1,4-GALACTOSYLTRANSFERASE-I COMPLEXED WITH ALPHA-LACTALBUMIN IN THE PRESENCE OF N-ACETYLGLUCOSAMINE, UDP AND MANGANESE
Biological Source:
Source Organism(s):
Mus musculus (Taxon ID: 10090)
Bos taurus (Taxon ID: 9913)
Bos taurus (Taxon ID: 9913)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.30 Å
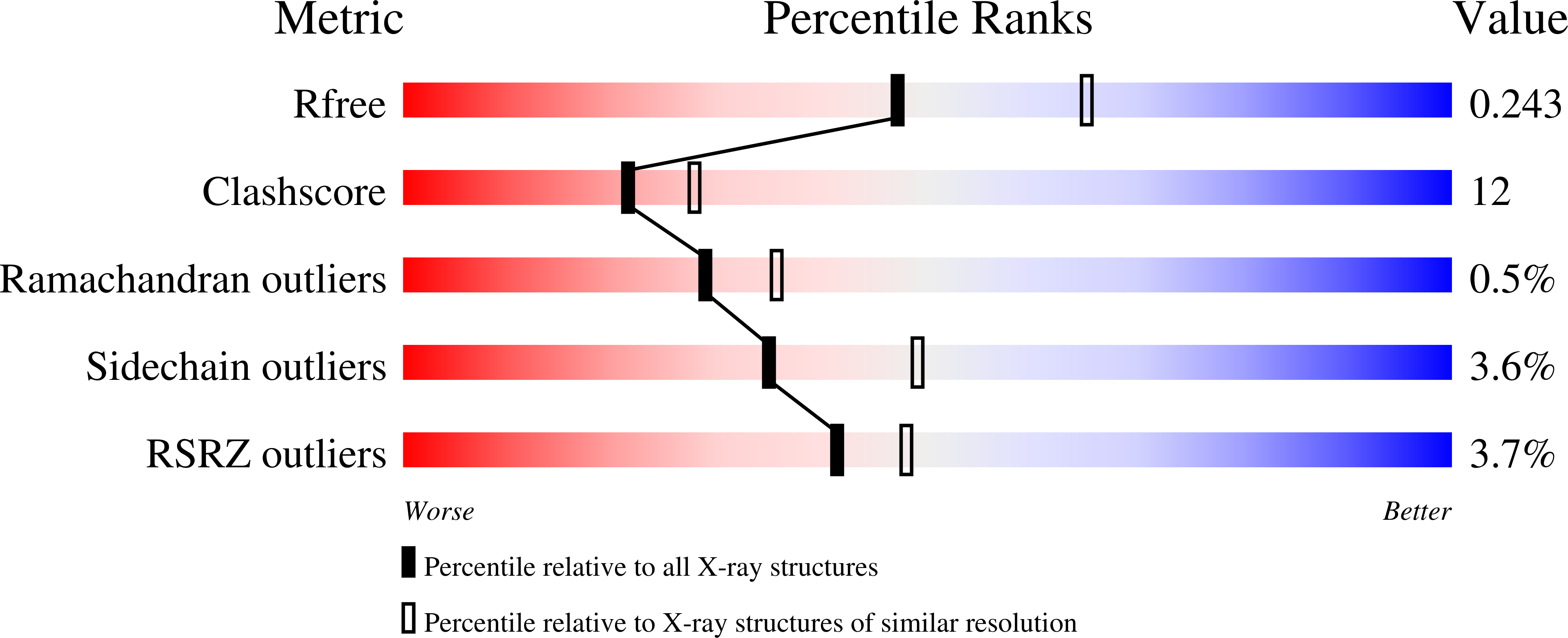
R-Value Free:
0.25
R-Value Work:
0.20
R-Value Observed:
0.20
Space Group:
P 1 21 1