Deposition Date
2003-05-28
Release Date
2003-11-11
Last Version Date
2023-08-16
Entry Detail
PDB ID:
1PGZ
Keywords:
Title:
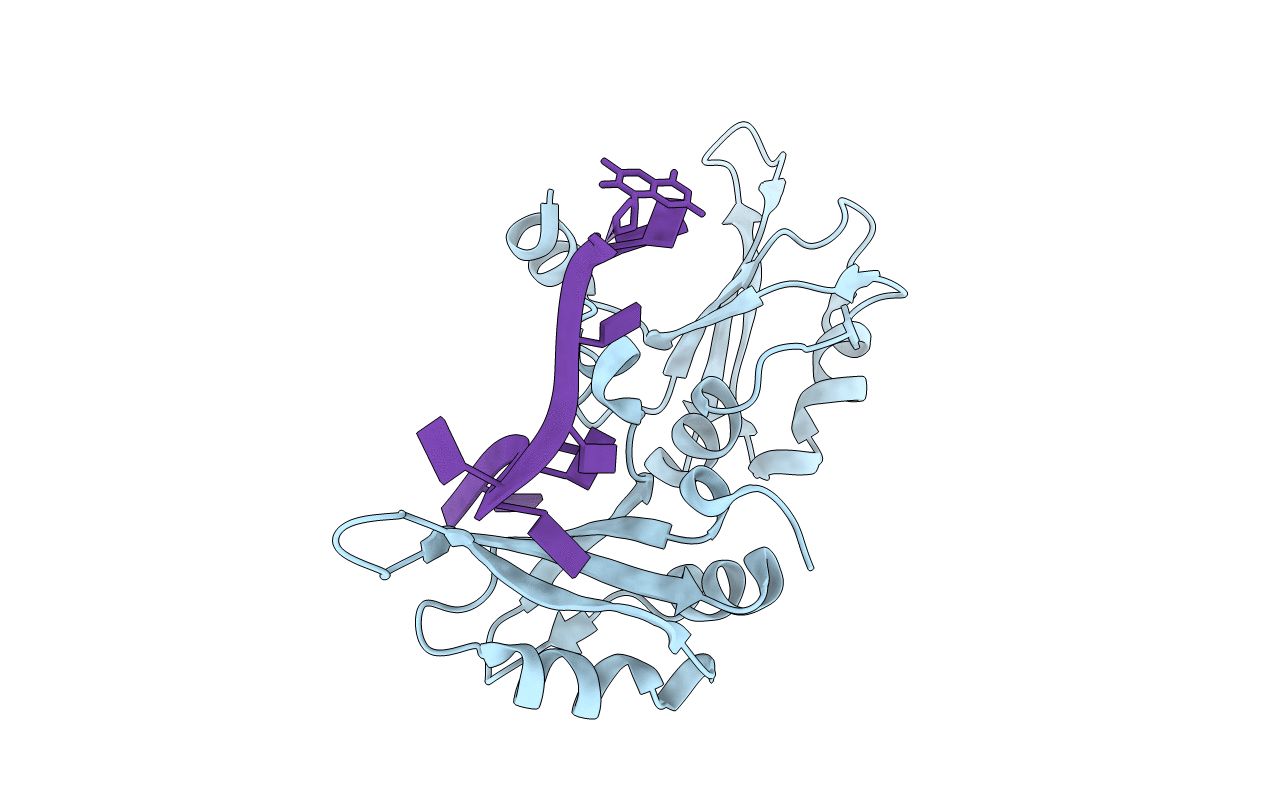
Crystal Structure of UP1 Complexed With d(TTAGGGTTAG(6-MI)G); A Human Telomeric Repeat Containing 6-methyl-8-(2-deoxy-beta-ribofuranosyl)isoxanthopteridine (6-MI)
Biological Source:
Source Organism:
Homo sapiens (Taxon ID: 9606)
Host Organism:
Method Details:
Experimental Method:
Resolution:
2.60 Å
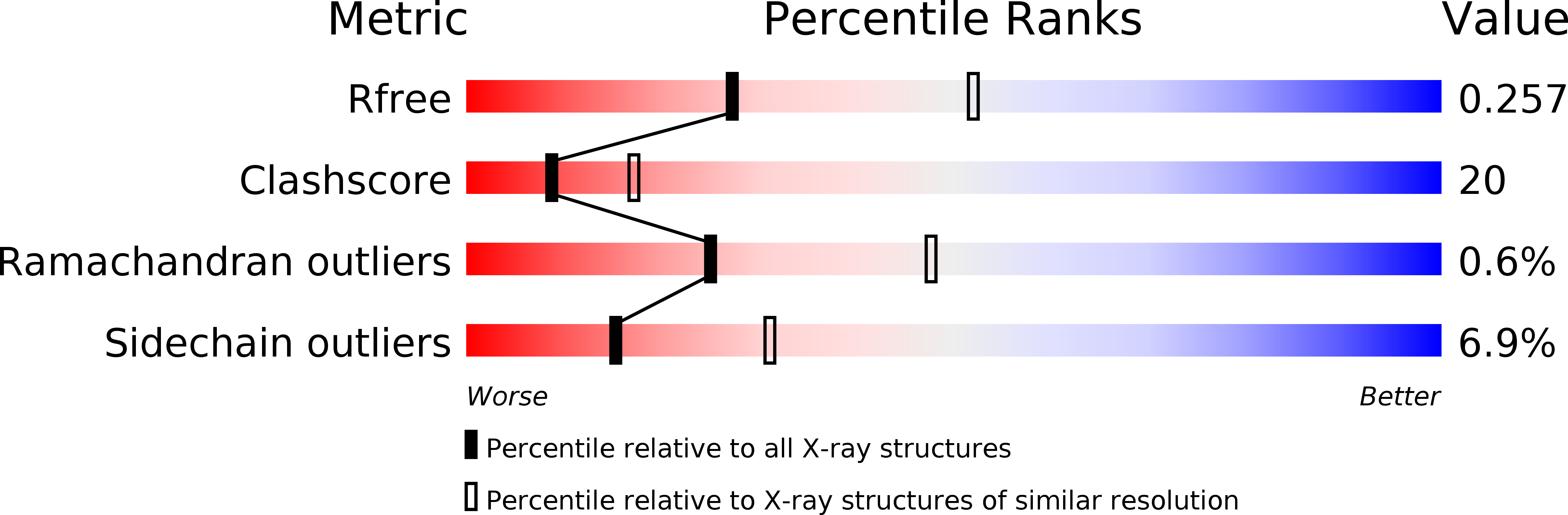
R-Value Free:
0.26
R-Value Work:
0.23
R-Value Observed:
0.23
Space Group:
P 43 21 2