Deposition Date
2003-03-03
Release Date
2003-03-25
Last Version Date
2024-10-23
Entry Detail
PDB ID:
1OO4
Keywords:
Title:
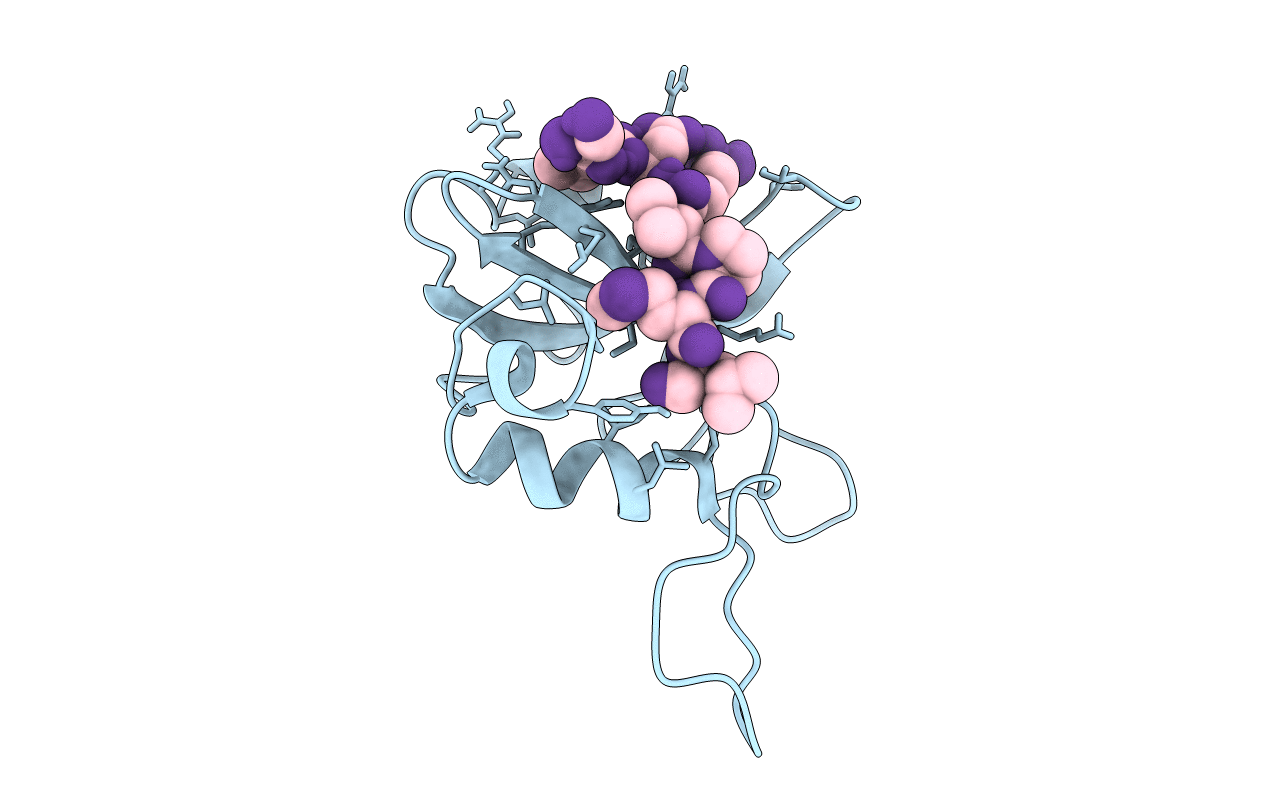
P395S mutant of the p85 regulatory subunit of the N-terminal src homology 2 domain of PI3-Kinase complexed to a peptide derived from PDGFr
Biological Source:
Source Organism(s):
Bos taurus (Taxon ID: 9913)
Expression System(s):
Method Details:
Experimental Method:
Conformers Submitted:
1