Abstact
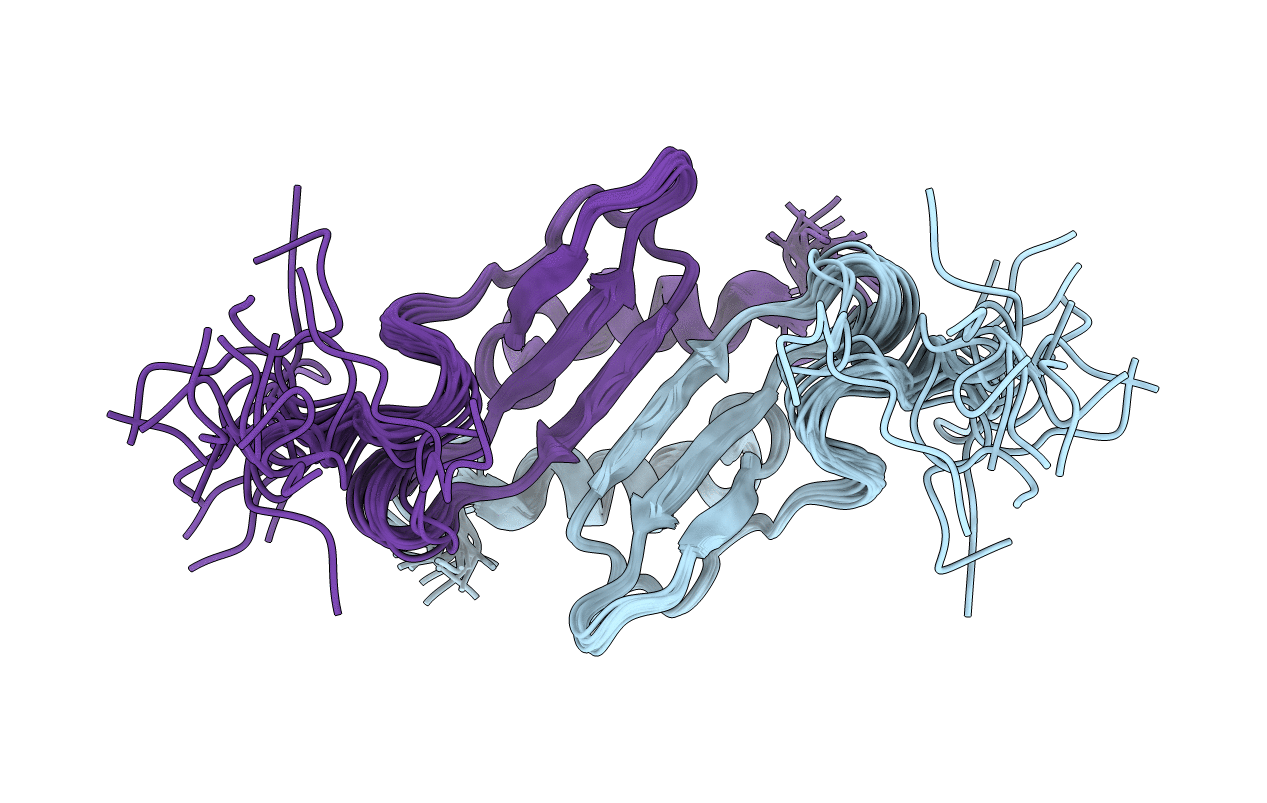
The solution structure of murine macrophage inflammatory protein-2 (MIP-2), a heparin-binding chemokine that is secreted in response to inflammatory stimuli, has been determined using two-dimensional homonuclear and heteronuclear NMR spectroscopy. Structure calculations were carried out by means of torsion-angle molecular dynamics using the program X-PLOR. The structure is based on a total of 2390 experimental restraints, comprising 2246 NOE-derived distance restraints, 44 distance restraints for 22 hydrogen bonds, and 100 torsion angle restraints. The structure is well-defined, with the backbone (N, Calpha, C) and heavy atom atomic rms distribution about the mean coordinates for residues 9-69 of the dimer being 0.57 +/- 0.16 A and 0.96 +/- 0.12 A, respectively. The N- and C-terminal residues (1-8 and 70-73, respectively) are disordered. The overall structure of the MIP-2 dimer is similar to that reported previously for the NMR structures of MGSA and IL-8 and consists of a six-stranded antiparallel beta-sheet (residue 25-29, 39-44, and 48-52) packed against two C-terminal antiparallel alpha-helices. A best fit superposition of the NMR structure of MIP-2 on the structures of MGSA, NAP-2, and the NMR and X-ray structures of IL-8 are 1.11, 1.02, 1.27, and 1.19 A, respectively, for the monomers, and 1.28, 1.10, 1.55, and 1.36 A, respectively, for the dimers (IL-8 residues 7-14 and 16-67, NAP-2 residues 25-84). At the tertiary level, the main differences between the MIP-2 solution structure and the IL-8, MGSA, and NAP-2 structures involve the N-terminal loop between residues 9-23 and the loops formed by residues 30-38 and residues 53-58. At the quaternary level, the difference between MIP-2 and IL-8, MGSA, or NAP-2 results from differing interhelical angles and separations.