Deposition Date
2001-06-05
Release Date
2001-08-22
Last Version Date
2024-11-13
Entry Detail
PDB ID:
1JBL
Keywords:
Title:
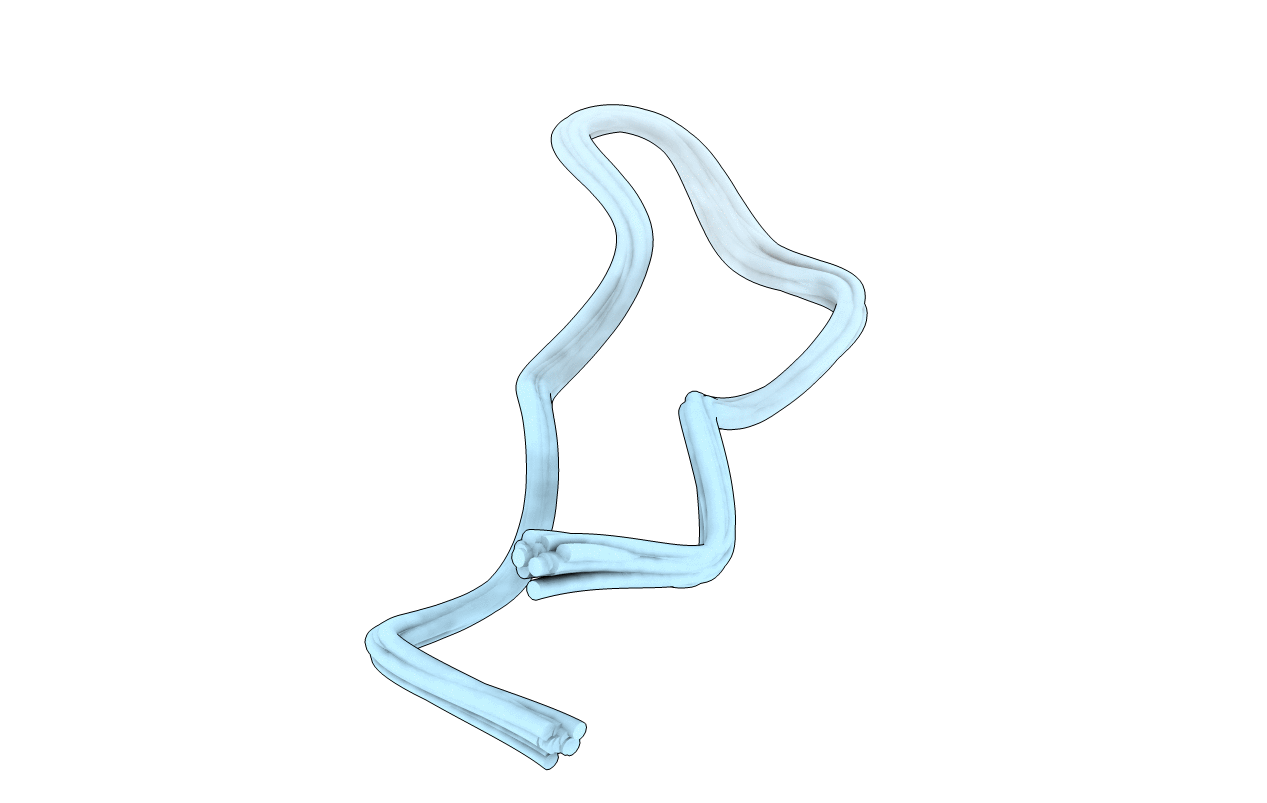
Solution structure of SFTI-1, A cyclic trypsin inhibitor from sunflower seeds
Biological Source:
Source Organism(s):
Helianthus annuus (Taxon ID: 4232)
Method Details:
Experimental Method:
Conformers Calculated:
50
Conformers Submitted:
20
Selection Criteria:
structures with the lowest energy