Abstact
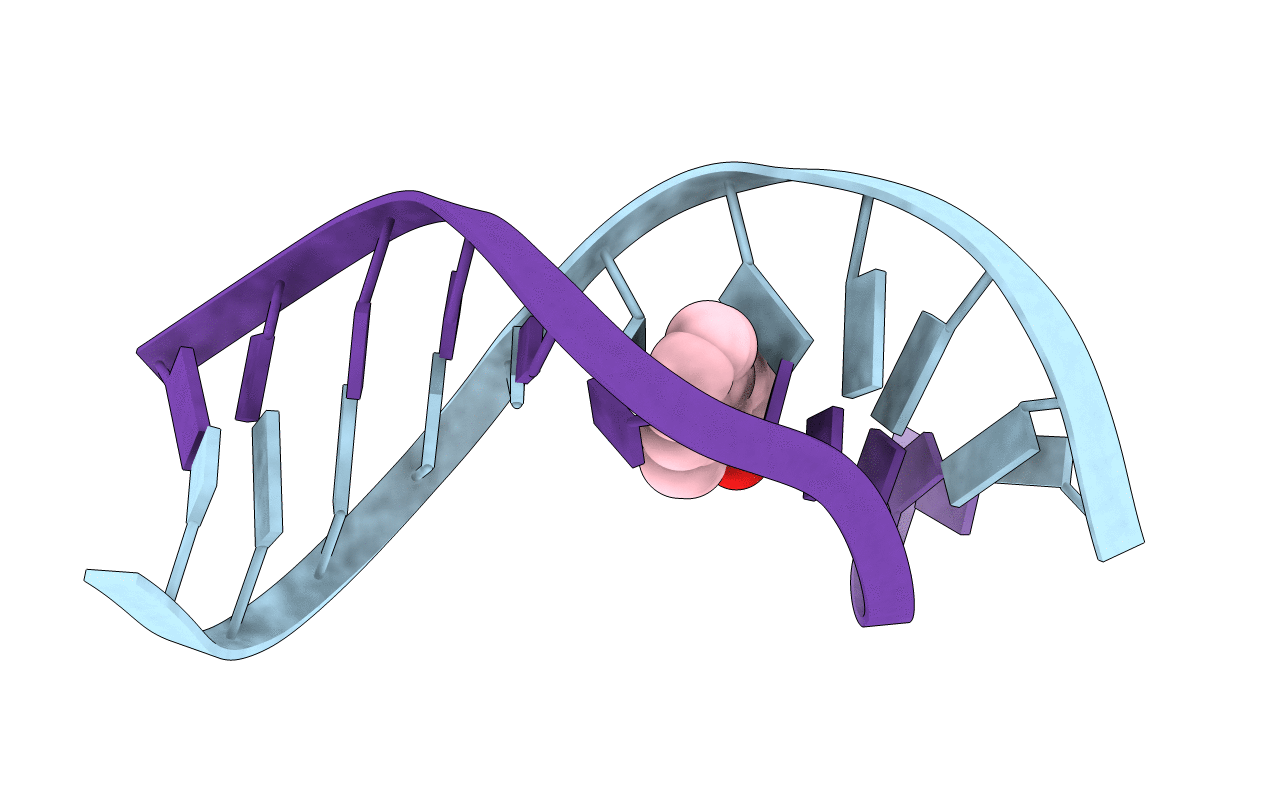
The structure of the bay region (1R,2S,3R,4S)-N6-[1-(1,2,3,4-tetrahydro-2,3,4-trihydroxybenz[a]anthracenyl)]-2'-deoxyadenosyl adduct at X(7) of 5'-d(CGGACAXGAAG)-3'.5'-d(CTTCTTGTCCG)-3', incorporating codons 60, 61 (underlined), and 62 of the human N-ras protooncogene, was determined by NMR. This was the bay region benz[a]anthracene RSRS (61,3) adduct. The BA moiety intercalated above the 5'-face of the modified base pair. NOE connectivities between imino protons were disrupted at T16 and T17. Large chemical shifts at the lesion site were consistent with ring current shielding arising from the BA moiety. A large chemical shift dispersion was observed for the BA aromatic protons. An increased rise of 8.17 A was observed between base pairs A6 x T17 and X7 x T(16). The PAH moiety stacked with the purine ring of A6, the 5'-neighbor nucleotide. This resulted in buckling of the 5'-neighbor A6 x T17 base pair, evidenced by exchange broadening for the T17 imino resonance. It also interrupted sequential NOE connectivities between nucleotides C5 and A6. The A6 deoxyribose ring showed an increased percentage of the C3'-endo conformation. This differed from the bay region BA RSRS (61,2) adduct, in which the lesion was located at position X6 [Li, Z., Mao, H., Kim, H.-Y., Tamura, P. J., Harris, C. M., Harris, T. M., and Stone, M. P. (1999) Biochemistry 38, 2969-2981], but was similar to the benzo[a]pyrene BP SRSR (61,3) adduct [Zegar I. S., Chary, P., Jabil, R. J., Tamura, P. J., Johansen, T. N., Lloyd, R. S., Harris, C. M., Harris, T. M., and Stone, M. P. (1998) Biochemistry 37, 16516-16528]. The altered sugar pseudorotation at A6 appears to be common to both bay region BA RSRS (61,3) and BP SRSR (61,3) adducts. It could not be discerned if the C3'-endo conformation at A6 in the BA RSRS (61,3) adduct altered base pairing geometry at X7 x T16, as compared to the C2'-endo conformation. The structural studies suggest that the mutational spectrum of this adduct may be more complex than that of the BA RSRS (61,2) adduct.