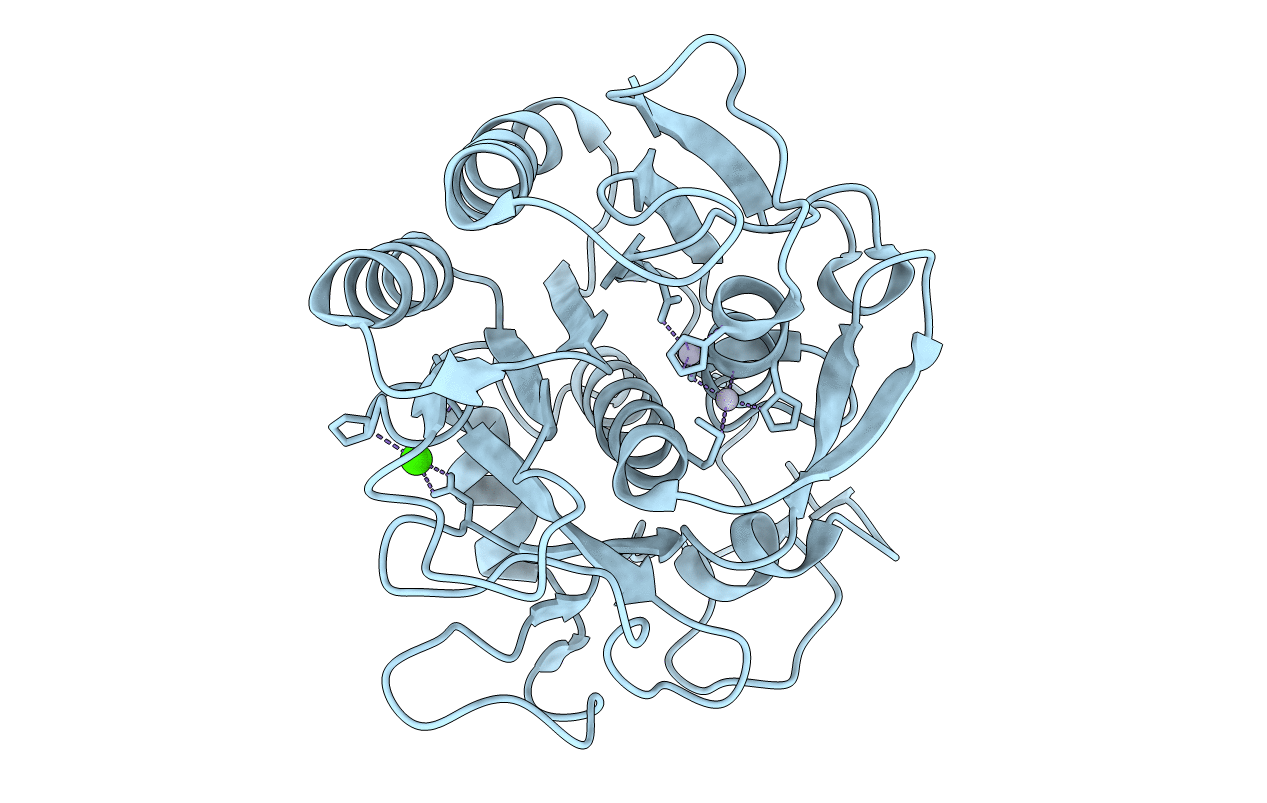
Deposition Date
2000-12-27
Release Date
2001-06-27
Last Version Date
2024-10-30
Entry Detail
PDB ID:
1HT3
Keywords:
Title:
MERCURY INDUCED MODIFICATIONS IN THE STEREOCHEMISTRY OF THE ACTIVE SITE THROUGH CYS-73 IN A SERINE PROTEASE: CRYSTAL STRUCTURE OF THE COMPLEX OF A PARTIALLY MODIFIED PROTEINASE K WITH MERCURY AT 1.8 A RESOLUTION
Biological Source:
Source Organism(s):
Engyodontium album (Taxon ID: 37998)
Method Details:
Experimental Method:
Resolution:
1.80 Å
R-Value Free:
0.22
R-Value Work:
0.16
R-Value Observed:
0.16
Space Group:
P 43 21 2


