Deposition Date
2000-10-21
Release Date
2000-11-01
Last Version Date
2024-02-07
Entry Detail
PDB ID:
1G2U
Keywords:
Title:
THE STRUCTURE OF THE MUTANT, A172V, OF 3-ISOPROPYLMALATE DEHYDROGENASE FROM THERMUS THERMOPHILUS HB8 : ITS THERMOSTABILITY AND STRUCTURE.
Biological Source:
Source Organism(s):
Thermus thermophilus (Taxon ID: 300852)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.10 Å
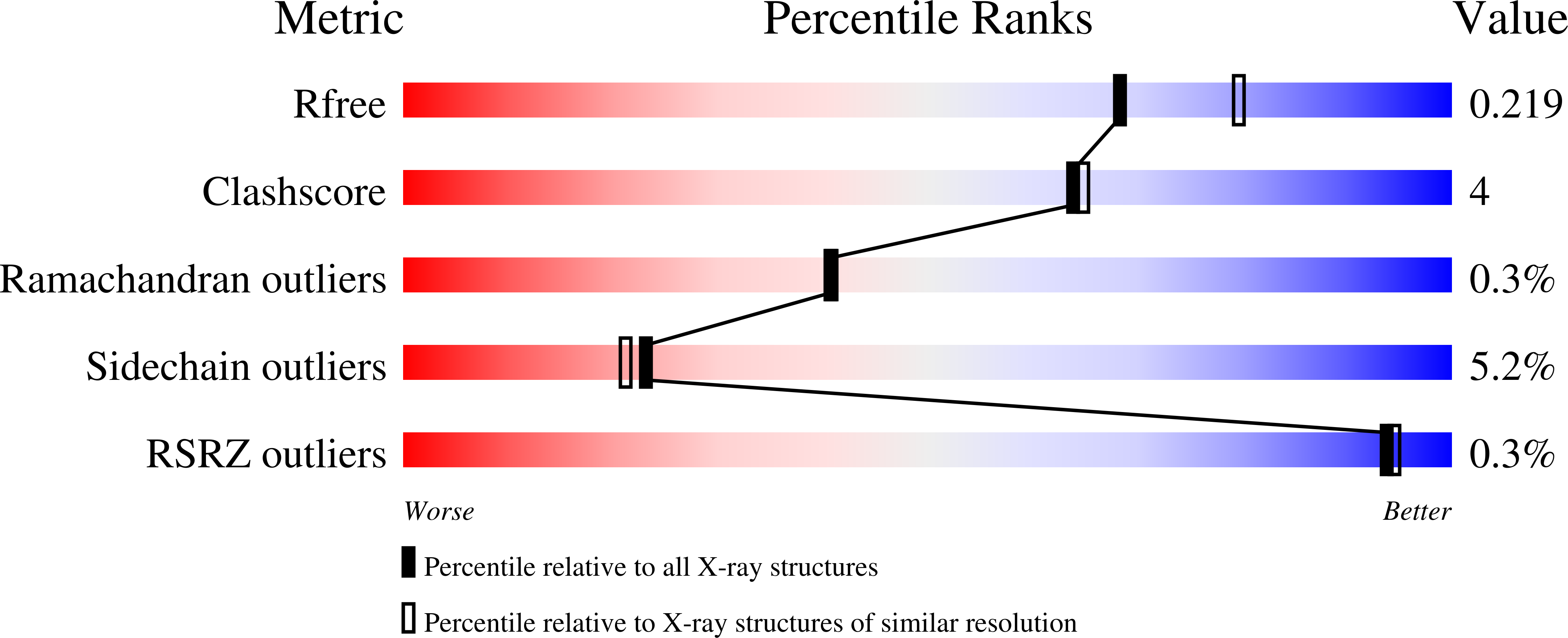
R-Value Free:
0.23
R-Value Work:
0.18
R-Value Observed:
0.18
Space Group:
P 32 2 1


