Deposition Date
1997-06-24
Release Date
1998-06-24
Last Version Date
2024-02-07
Entry Detail
PDB ID:
1FSZ
Keywords:
Title:
CRYSTAL STRUCTURE OF THE CELL-DIVISION PROTEIN FTSZ AT 2.8A RESOLUTION
Biological Source:
Source Organism(s):
Methanocaldococcus jannaschii (Taxon ID: 2190)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.80 Å
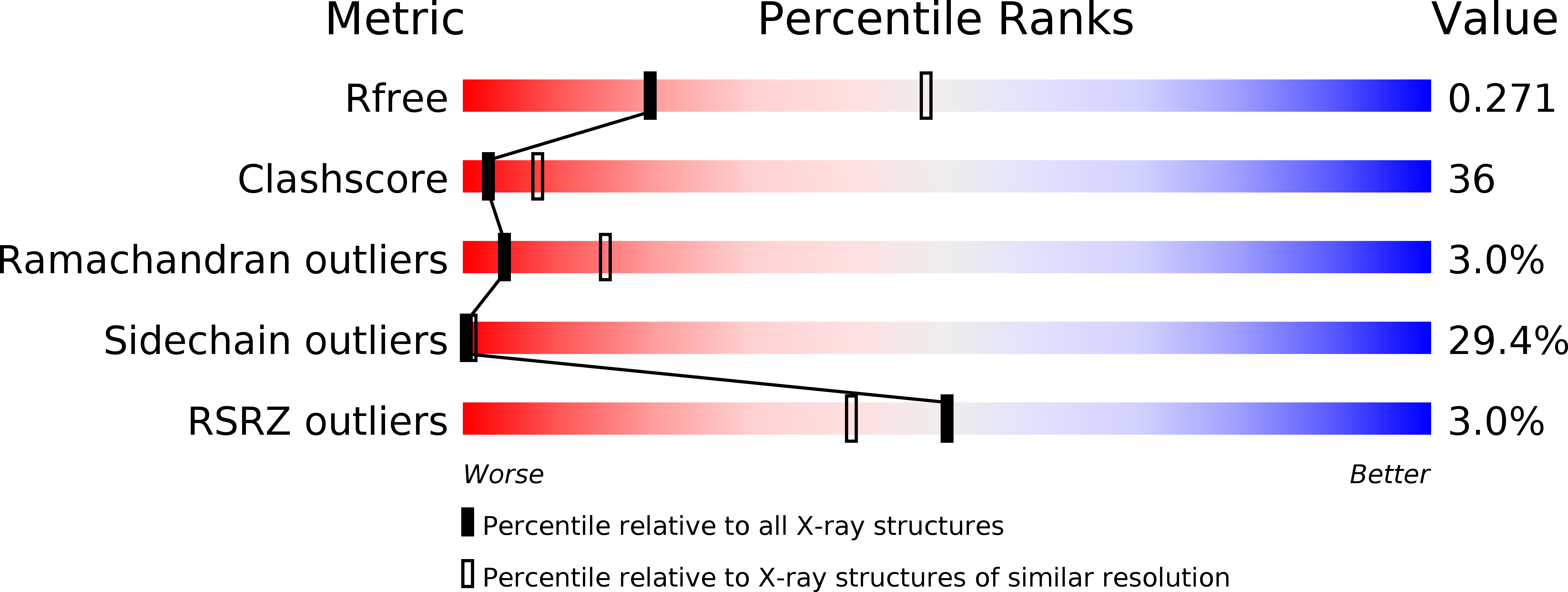
R-Value Free:
0.28
R-Value Work:
0.19
R-Value Observed:
0.19
Space Group:
I 21 3