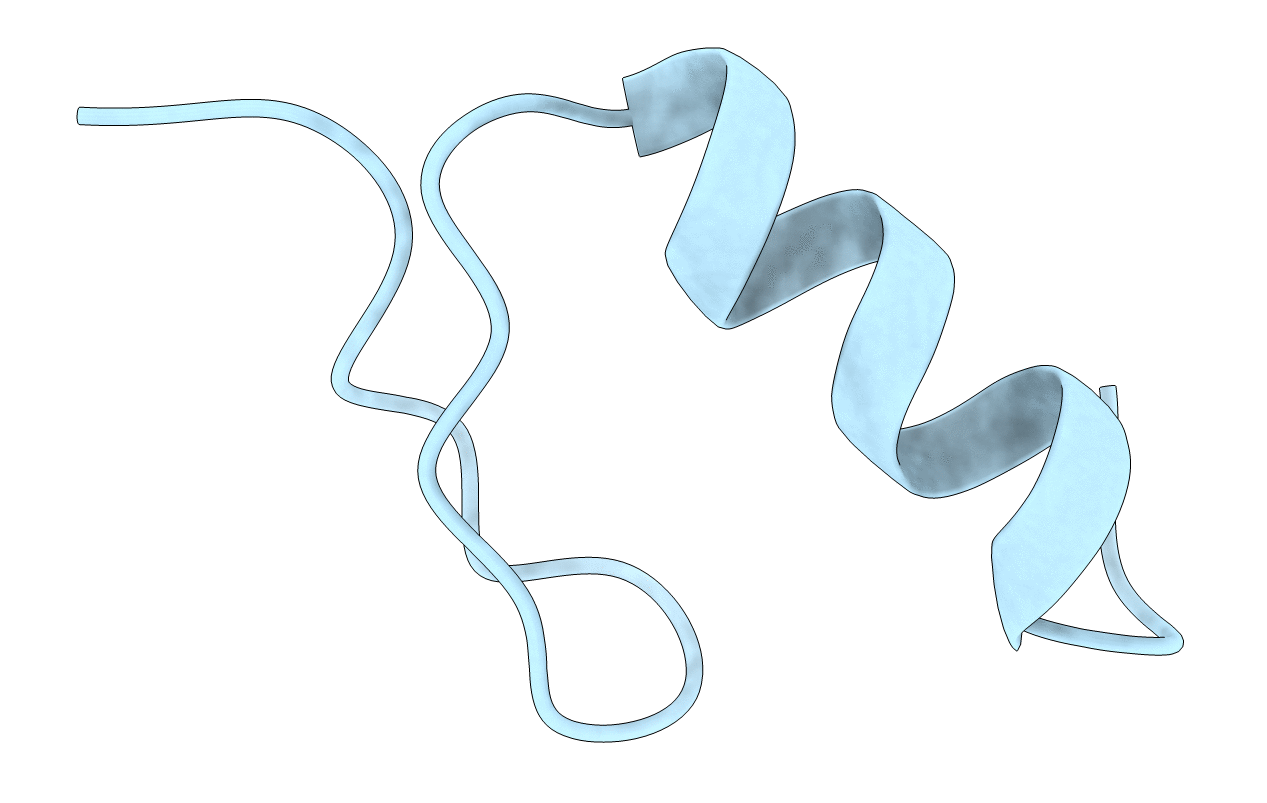
Deposition Date
1997-10-26
Release Date
1998-01-28
Last Version Date
2024-05-22
Entry Detail
PDB ID:
1FSV
Keywords:
Title:
FULL SEQUENCE DESIGN 1 (FSD-1) OF BETA BETA ALPHA MOTIF, NMR, MINIMIZED AVERAGE STRUCTURE
Biological Source:
Source Organism(s):
synthetic construct (Taxon ID: 32630)
Method Details:
Experimental Method:
Conformers Calculated:
100
Conformers Submitted:
1
Selection Criteria:
RESTRAINED MINIMIZED AVERAGE


