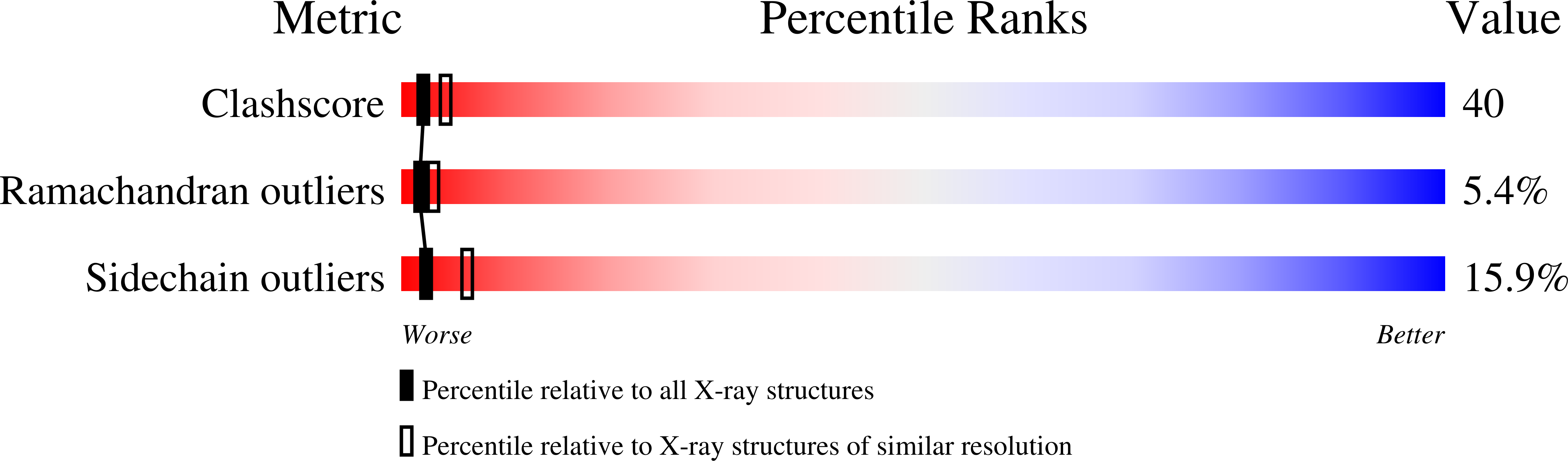
Deposition Date
2000-05-12
Release Date
2000-11-22
Last Version Date
2024-02-07
Entry Detail
PDB ID:
1EZZ
Keywords:
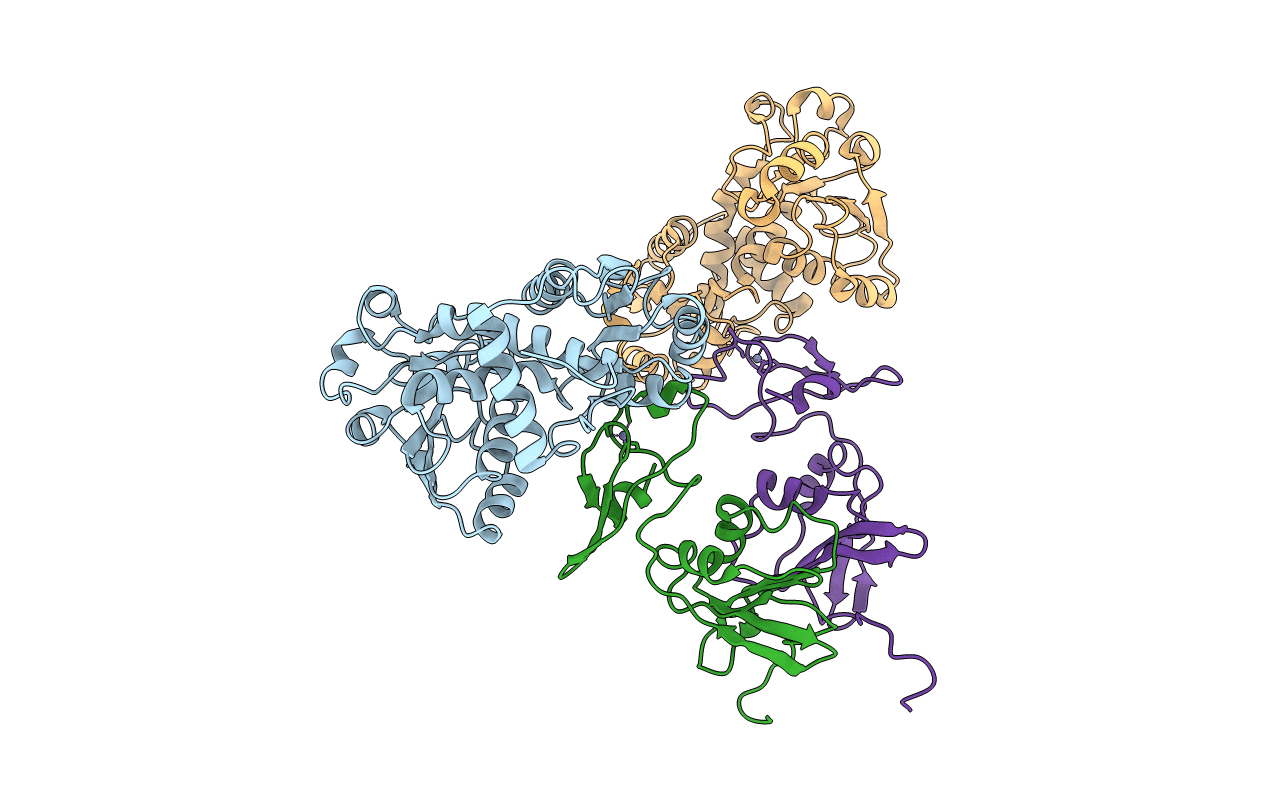
Title:
CRYSTAL STRUCTURE OF E. COLI ASPARTATE TRANSCARBAMOYLASE P268A MUTANT IN THE T-STATE
Biological Source:
Source Organism:
Escherichia coli (Taxon ID: 562)
Host Organism:
Method Details:
Experimental Method:
Resolution:
2.70 Å
R-Value Free:
0.24
R-Value Work:
0.18
Space Group:
H 3