Deposition Date
1992-03-31
Release Date
1993-07-15
Last Version Date
2024-02-07
Entry Detail
PDB ID:
1D67
Keywords:
Title:
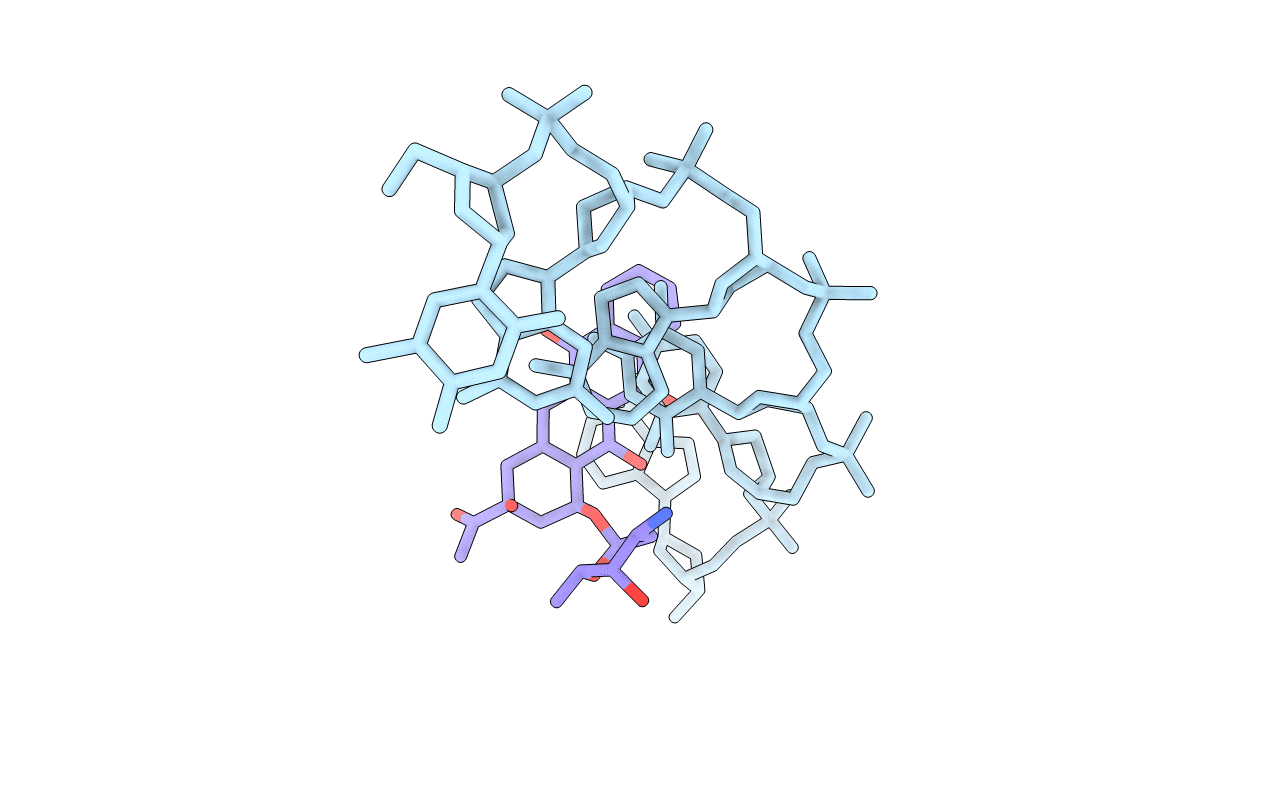
THE MOLECULAR STRUCTURE OF AN IDARUBICIN-D(TGATCA) COMPLEX AT HIGH RESOLUTION
Method Details:
Experimental Method:
Resolution:
1.60 Å
R-Value Observed:
0.22
Space Group:
P 41 21 2