Deposition Date
1999-10-04
Release Date
2000-10-11
Last Version Date
2021-11-03
Entry Detail
PDB ID:
1D4L
Keywords:
Title:
HIV-1 PROTEASE COMPLEXED WITH A MACROCYCLIC PEPTIDOMIMETIC INHIBITOR
Method Details:
Experimental Method:
Resolution:
1.75 Å
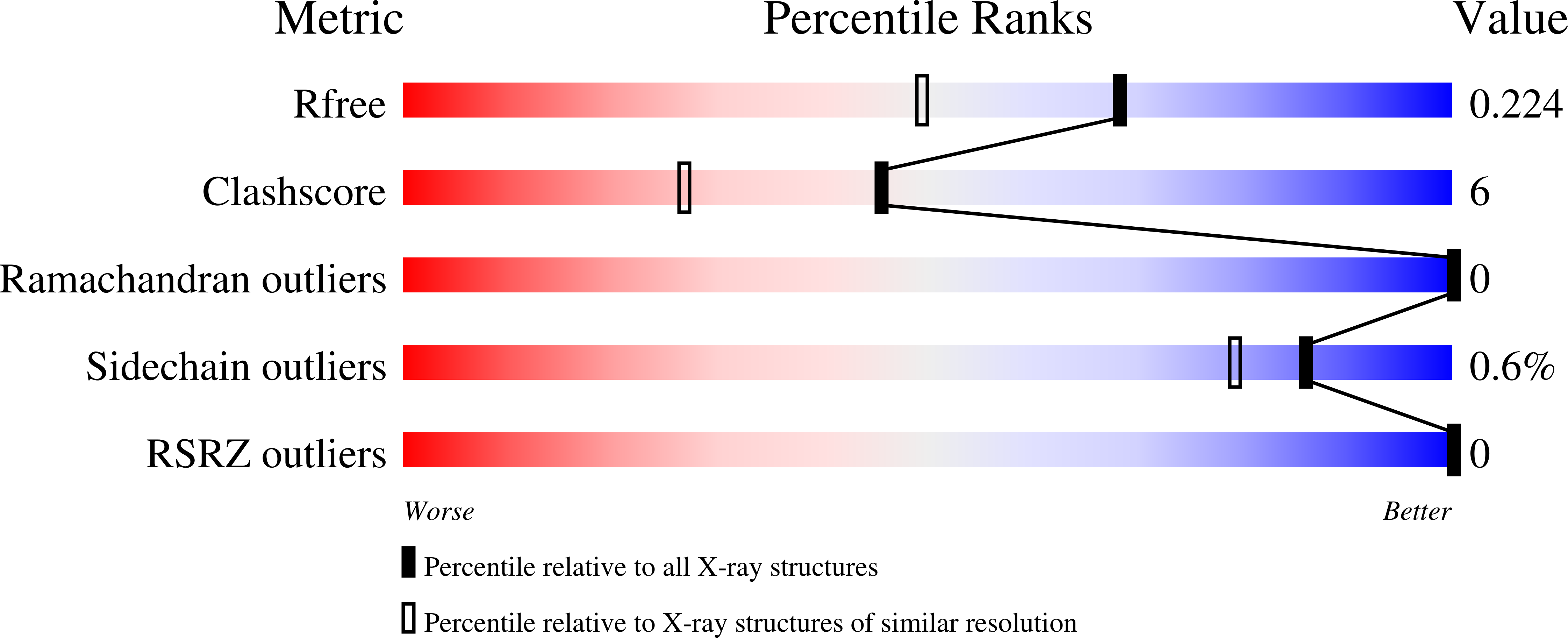
R-Value Free:
0.23
R-Value Work:
0.18
R-Value Observed:
0.18
Space Group:
P 21 21 21