Deposition Date
1999-08-12
Release Date
1999-08-31
Last Version Date
2024-02-07
Entry Detail
PDB ID:
1CQX
Keywords:
Title:
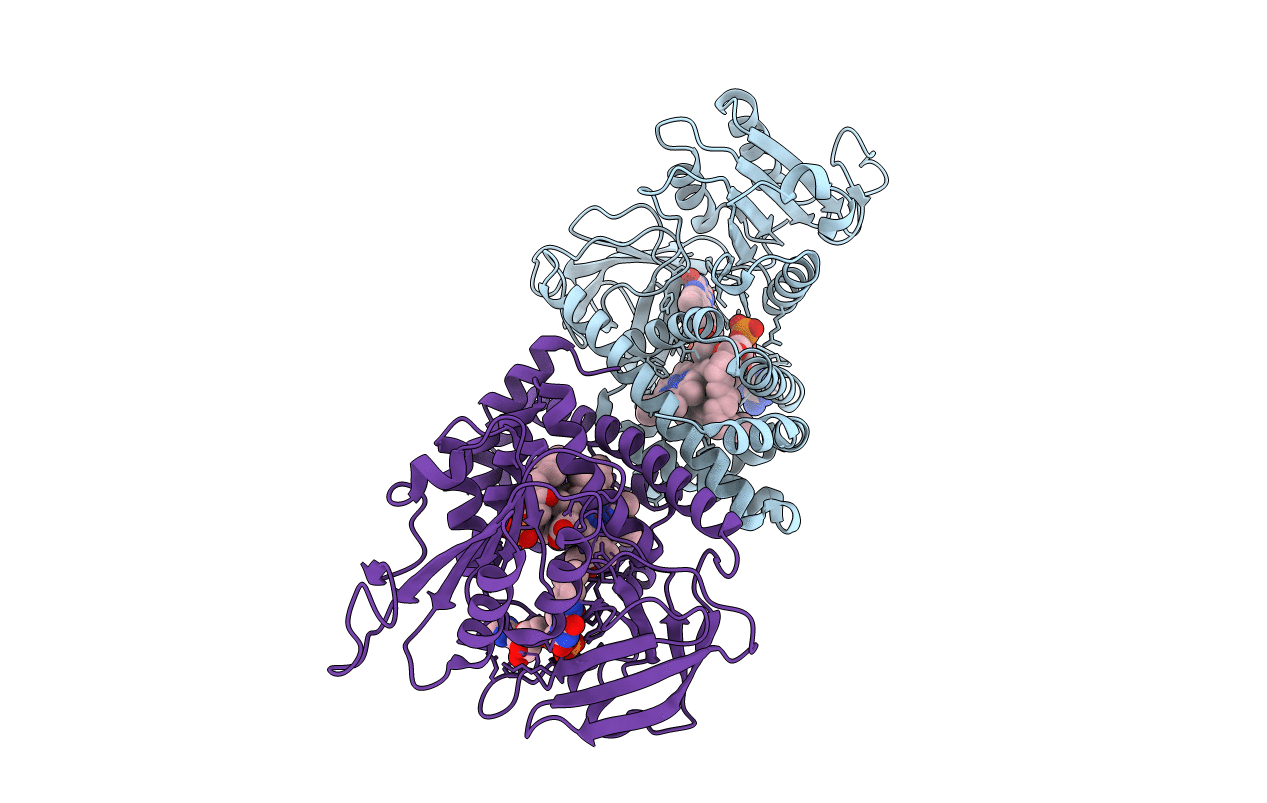
Crystal structure of the flavohemoglobin from Alcaligenes eutrophus at 1.75 A resolution
Biological Source:
Source Organism(s):
Cupriavidus necator (Taxon ID: 106590)
Method Details:
Experimental Method:
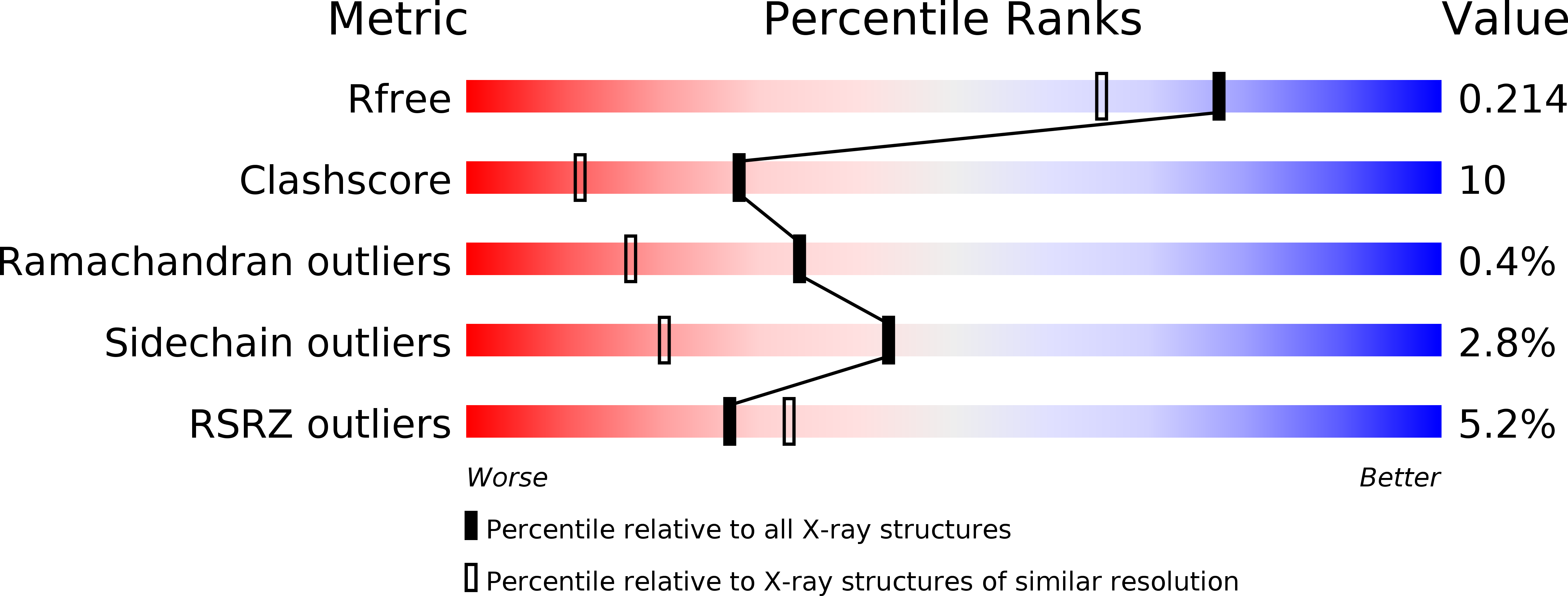
Resolution:
1.75 Å
R-Value Free:
0.21
R-Value Work:
0.18
Space Group:
P 1 21 1


