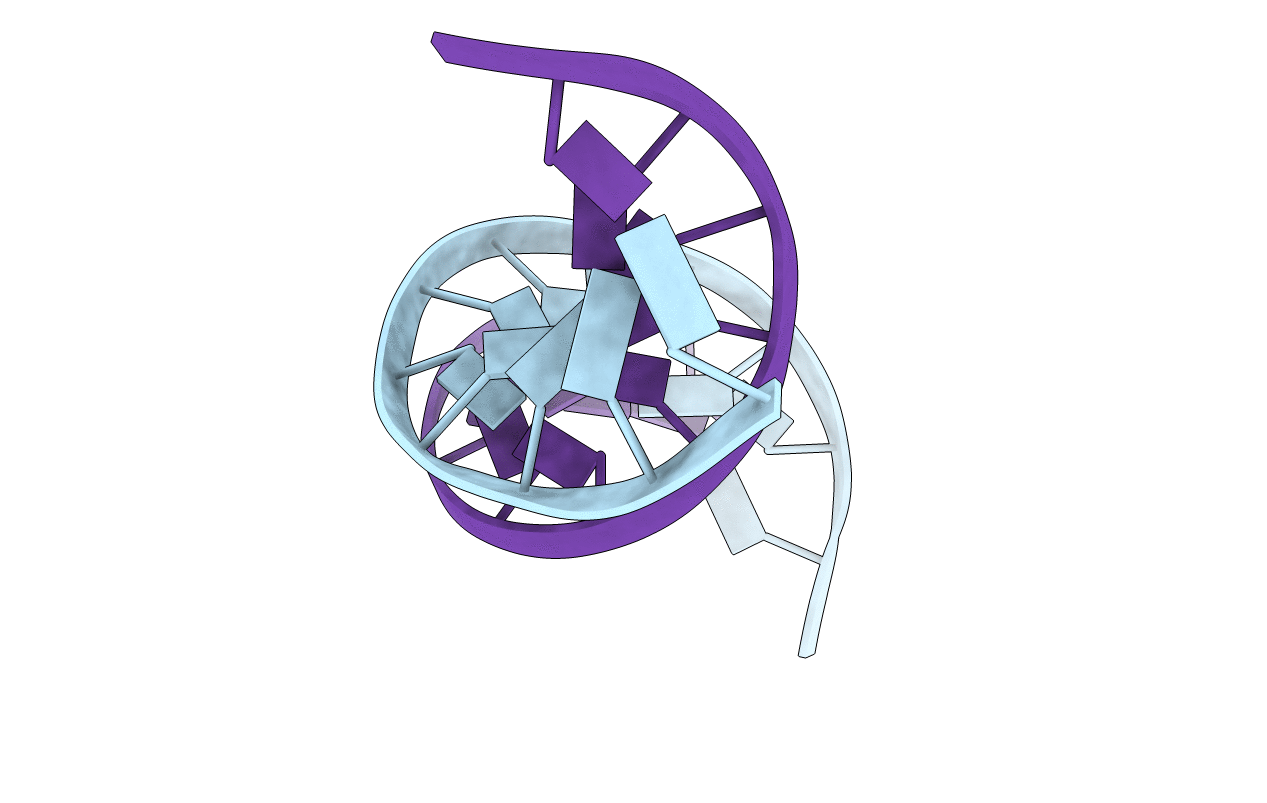
Deposition Date
1999-05-26
Release Date
1999-06-09
Last Version Date
2023-12-27
Entry Detail
PDB ID:
1COC
Keywords:
Title:
SOLUTION-STATE STRUCTURE OF A DNA DODECAMER DUPLEX CONTAINING A CIS-SYN THYMINE CYCLOBUTANE DIMER.
Method Details:
Experimental Method:
Conformers Submitted:
1


