Deposition Date
1995-08-30
Release Date
1996-11-08
Last Version Date
2024-05-22
Entry Detail
Method Details:
Experimental Method:
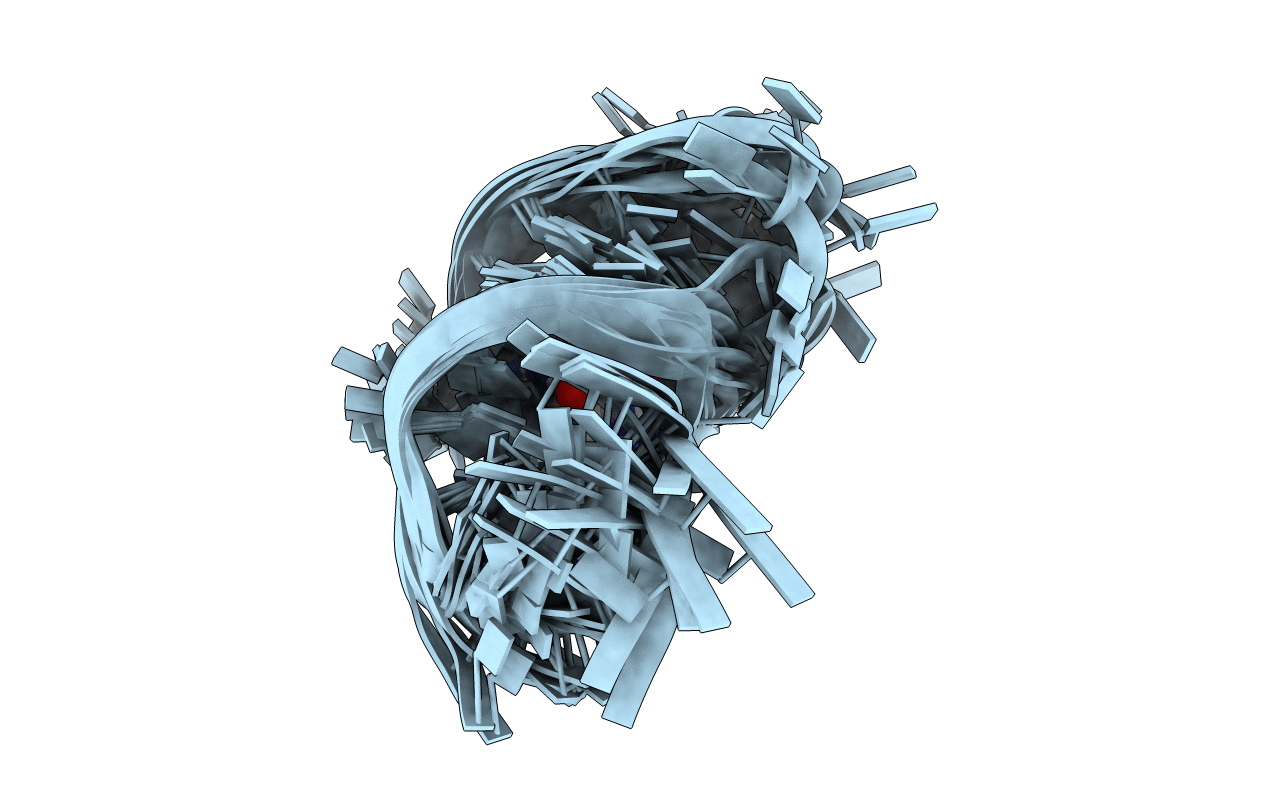
Conformers Submitted:
20