Deposition Date
1998-02-06
Release Date
1998-04-29
Last Version Date
2024-10-16
Entry Detail
PDB ID:
1A4W
Keywords:
Title:
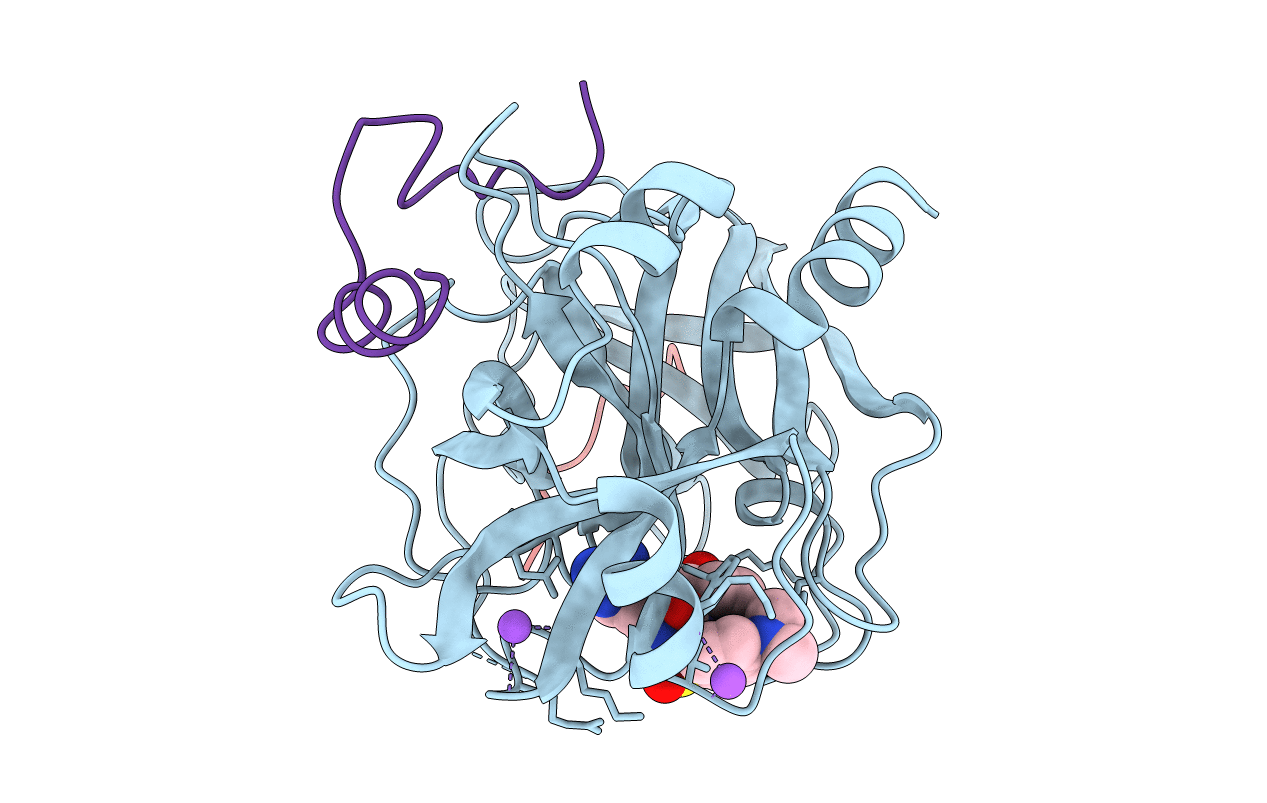
CRYSTAL STRUCTURES OF THROMBIN WITH THIAZOLE-CONTAINING INHIBITORS: PROBES OF THE S1' BINDING SITE
Biological Source:
Source Organism(s):
Homo sapiens (Taxon ID: 9606)
Method Details: