Search Count: 103
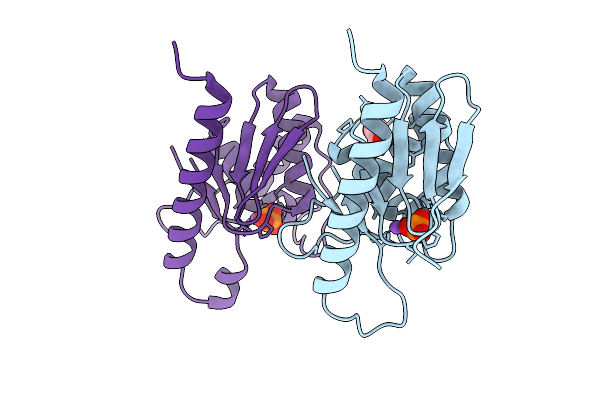 |
High Resolution Structure Of Cytidine Deaminase T6S Toxin From Pseudomonas Syringae
Organism: Pseudomonas syringae
Method: X-RAY DIFFRACTION Release Date: 2025-11-12 Classification: TOXIN Ligands: ZN, PO4, EDO, NA |
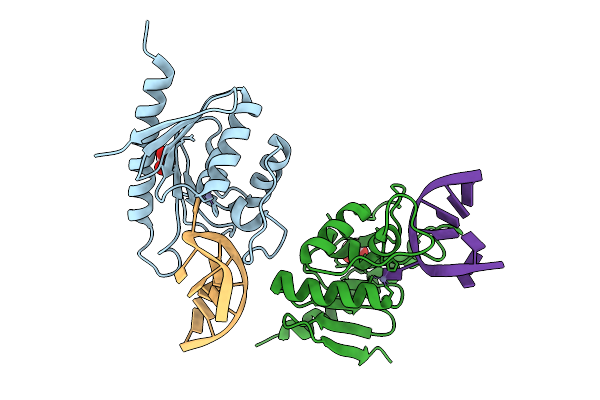 |
Cytidine Deaminase T6S Toxin From Pseudomonas Syringae Complexed With Substrate Dna
Organism: Pseudomonas syringae
Method: X-RAY DIFFRACTION Release Date: 2025-11-12 Classification: TOXIN/DNA Ligands: ZN, EDO |
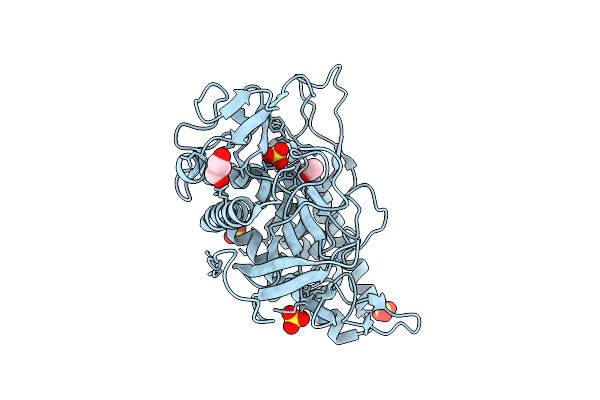 |
Crystal Structure Of The Catalytic Domain From The Bont-Like Toxin Complex Pg1 Of Paeniclostridium Ghonii
Organism: Paraclostridium ghonii
Method: X-RAY DIFFRACTION Release Date: 2025-10-15 Classification: TOXIN Ligands: ZN, SO4, ACT, GOL |
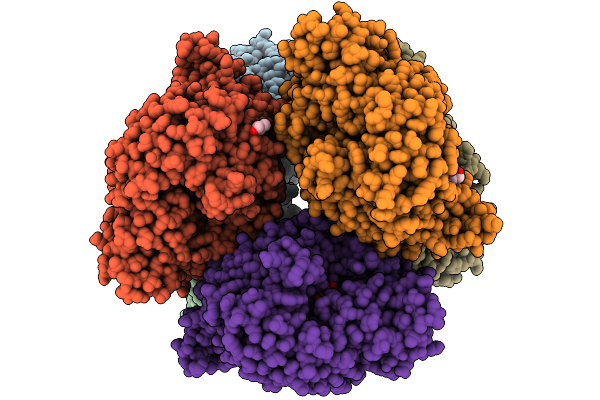 |
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GLC, GOL, NA, G1P |
 |
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Release Date: 2025-09-10 Classification: STRUCTURAL PROTEIN Ligands: GOL, GLC, PEG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-09-03 Classification: TRANSPORT PROTEIN Ligands: URE, PLM |
 |
Structural Basis Of The Bifunctionality Of M. Salinexigens Zyf650T Glucosylglycerol Phosphorylase In Glucosylglycerol Catabolism
Organism: Marinobacter salinexigens
Method: X-RAY DIFFRACTION Release Date: 2025-09-03 Classification: STRUCTURAL PROTEIN Ligands: PEG, TRS, PO4, SO4 |
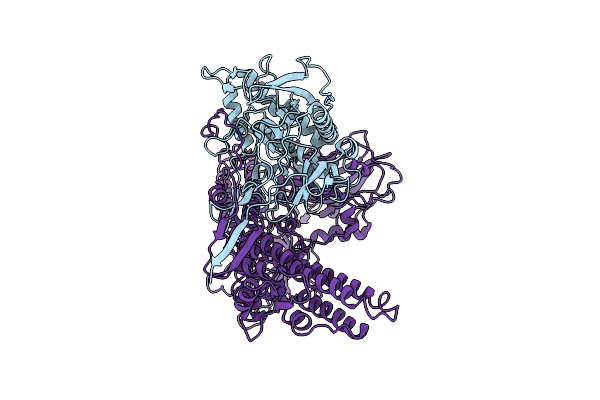 |
Cryo-Em Structure And Molecular Assembly Of The Bont-Like Toxin Pg1 Complex From Paeniclostridium Ghonii
Organism: Paraclostridium ghonii
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2025-03-26 Classification: TOXIN |
 |
Structural Basis Of Elongation Factor Tu In Regulating Photoinhibition In Synechococcus Sp. Pcc 7942.
Organism: Synechococcus elongatus (strain atcc 33912 / pcc 7942 / fachb-805)
Method: X-RAY DIFFRACTION Release Date: 2025-03-19 Classification: PHOTOSYNTHESIS Ligands: GOL, GDP, BME, MG |
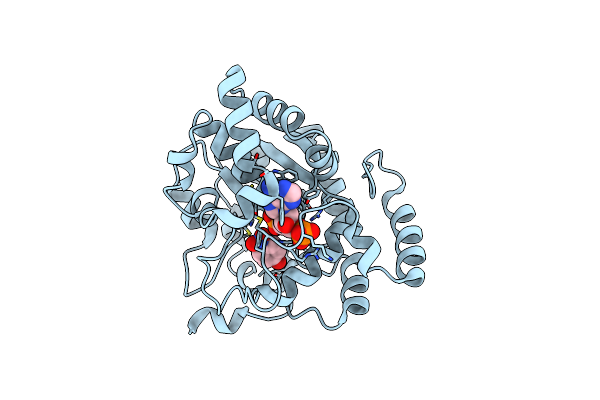 |
Crystal Structure Of A Sulfotransferase S4 From In Complex With Pap And Pnp
Organism: Bambusicola thoracicus
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2025-01-15 Classification: TRANSFERASE Ligands: NPO, A3P |
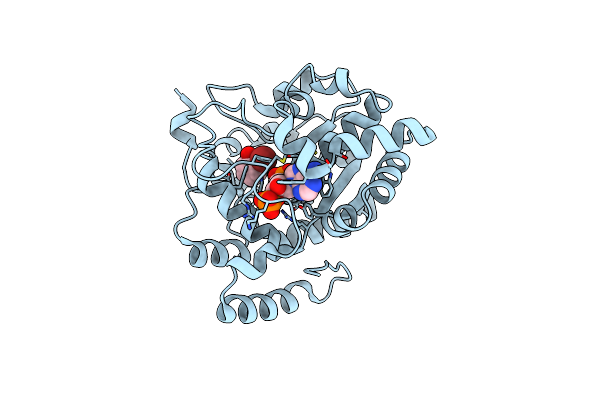 |
Crystal Structure Of A Sulfotransferase S4 In Complex With Pap And 2-Phe-Br
Organism: Bambusicola thoracicus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-12-25 Classification: TRANSFERASE Ligands: A3P, 2BR |
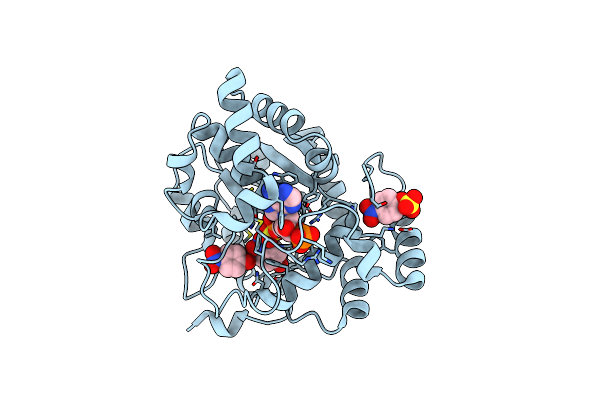 |
Organism: Bambusicola thoracicus
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2024-12-25 Classification: TRANSFERASE Ligands: NPO, PPS, 4NS |
 |
Organism: Bambusicola thoracicus
Method: X-RAY DIFFRACTION Resolution:2.44 Å Release Date: 2024-12-18 Classification: TRANSFERASE Ligands: A3P |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.87 Å Release Date: 2024-12-04 Classification: IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2024-12-04 Classification: IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2024-08-21 Classification: HYDROLASE Ligands: ZN, EDO, CL, NA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2024-08-21 Classification: HYDROLASE Ligands: ZN, EDO, CL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2024-08-21 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2024-08-21 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2024-08-21 Classification: HYDROLASE Ligands: ZN, EDO |

