Search Count: 6,072
All
Selected
 |
Composite Map For Cryo-Em Structure Of Dnmt3A2-Dnmt3B3 Tetramer Bound To 167H3K36Me2-Nucleosome
Organism: Xenopus laevis, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: DNA BINDING PROTEIN Ligands: ZN, SAO |
 |
Organism: Homo sapiens, Xenopus laevis, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: GENE REGULATION/DNA |
 |
Organism: Xenopus laevis, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: GENE REGULATION/DNA |
 |
Cryo-Em Structure Of Dnmt3A2/3B3 In Complex With H3K36Me2 Di-Nucleosome With Eight Base Pair Linker
Organism: Xenopus laevis, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: DNA BINDING PROTEIN Ligands: ZN, SAH |
 |
Cryo-Em Structure Of The Xenopus Laevis Mitotic Centromere-Associated Kinesin (Mcak) Bound To The Paclitaxel-Stabilized Microtubule
Organism: Xenopus laevis, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CELL CYCLE Ligands: GTP, MG, GDP, TA1 |
 |
Cryo-Em Structure Of The Xenopus Laevis Mitotic Centromere-Associated Kinesin (Mcak) Bound To The Paclitaxel-Stabilized Microtubule
Organism: Xenopus laevis, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CELL CYCLE Ligands: GTP, MG, GDP, TA1 |
 |
Organism: Homo sapiens, Xenopus laevis
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
 |
Cryo-Em Structure Of Ro60/La/Minimal Misfolded Pre-5S Rrna Complex With Fab, Composite Map
Organism: Homo sapiens, Xenopus laevis
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-02-04 Classification: RNA BINDING PROTEIN/RNA Ligands: MG |
 |
Structural Characterisation Of Chromatin Remodelling Intermediates Supports Linker Dna Dependent Product Inhibition As A Mechanism For Nucleosome Spacing.
Organism: Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2026-02-04 Classification: GENE REGULATION |
 |
Cryo-Em Structure Of The Xenopus Laevis Mitotic Centromere-Associated Kinesin (Mcak) Bound To The Paclitaxel-Stabilized Microtubule
Organism: Xenopus laevis, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: CELL CYCLE Ligands: MG, GTP, GDP, TA1 |
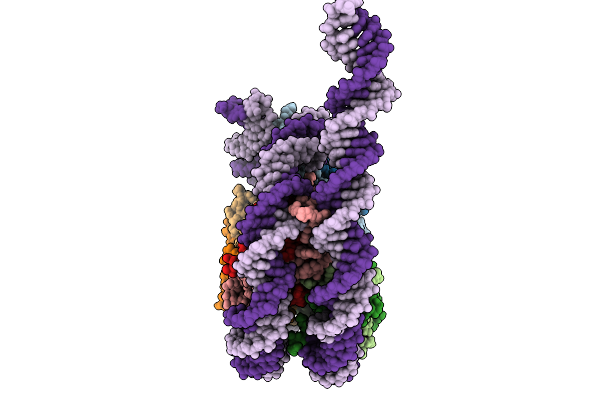 |
Structural Characterisation Of Chromatin Remodelling Intermediates Supports Linker Dna Dependent Product Inhibition As A Mechanism For Nucleosome Spacing.
Organism: Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:4.20 Å Release Date: 2026-01-28 Classification: GENE REGULATION |
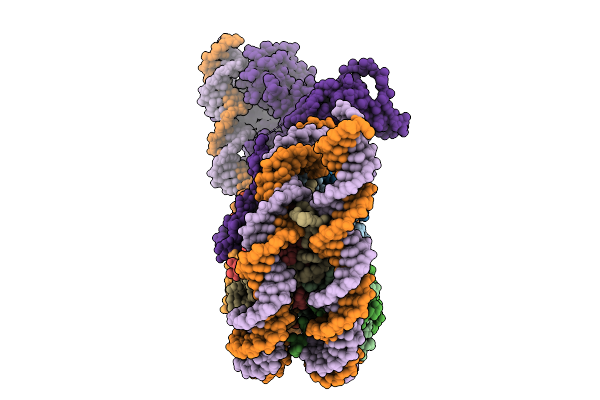 |
Structural Characterisation Of Chromatin Remodelling Intermediates Supports Linker Dna Dependent Product Inhibition As A Mechanism For Nucleosome Spacing.
Organism: Xenopus laevis, Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2026-01-28 Classification: GENE REGULATION |
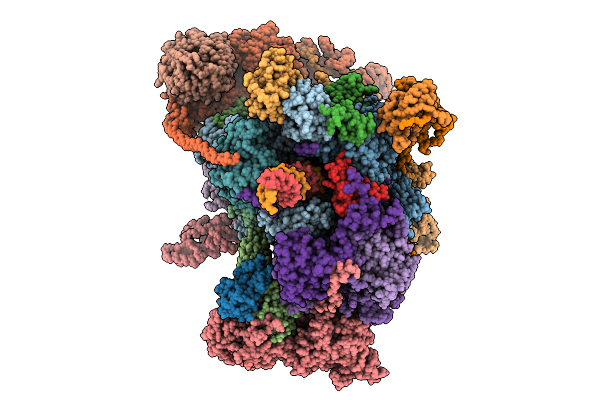 |
Pol Ii-Dsif-Spt6-Paf1C-Tfiis-Iws1-Elof1-Ledgf-Nucleosome Activated Elongation Complex Composite Map T
Organism: Homo sapiens, Xenopus laevis, Synthetic construct, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: TRANSCRIPTION Ligands: ZN, MG |
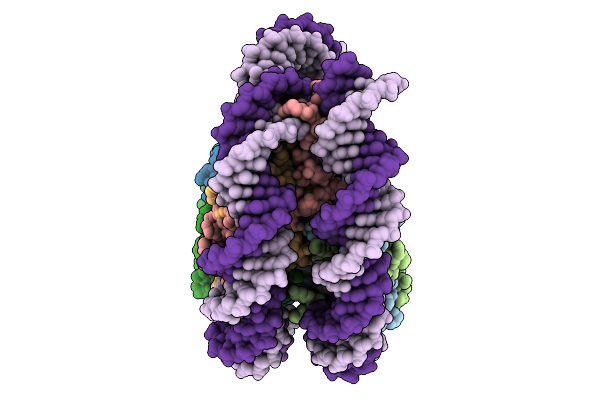 |
Organism: Xenopus laevis laevis, Xenopus laevis
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2026-01-21 Classification: TRANSLOCASE |
 |
Organism: Xenopus laevis, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-14 Classification: TRANSPORT PROTEIN |
 |
Organism: Xenopus laevis, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:7.50 Å Release Date: 2026-01-14 Classification: TRANSPORT PROTEIN |
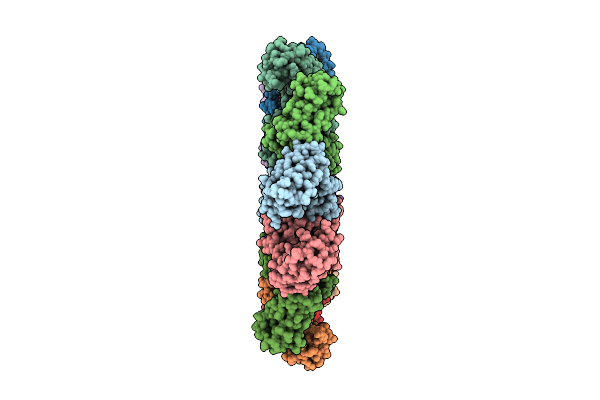 |
Organism: Xenopus laevis
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2025-12-17 Classification: IMMUNE SYSTEM |
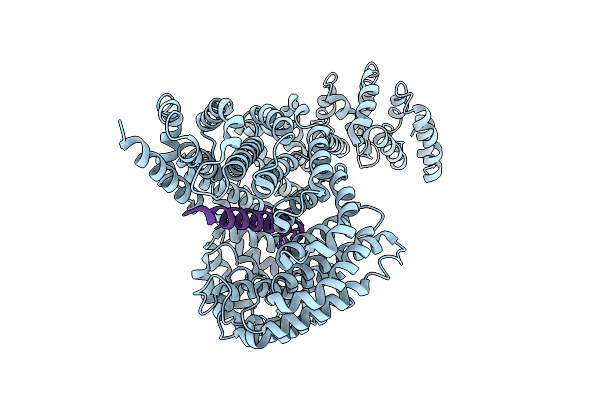 |
Organism: Homo sapiens, Xenopus laevis
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2025-12-10 Classification: TRANSPORT PROTEIN |
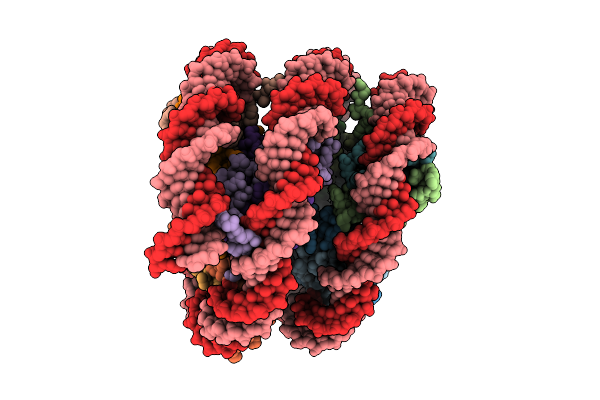 |
Organism: Xenopus laevis, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:4.94 Å Release Date: 2025-12-10 Classification: GENE REGULATION |
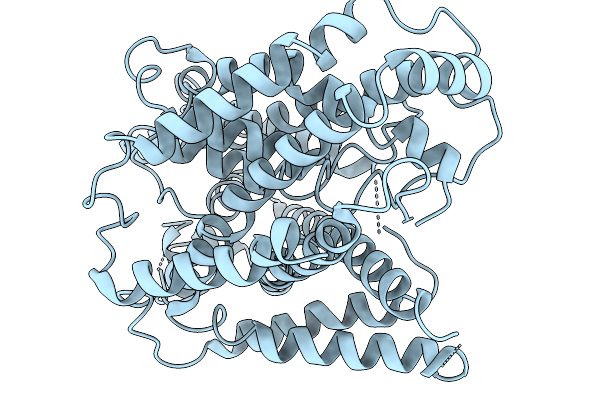 |
Organism: Xenopus tropicalis
Method: ELECTRON MICROSCOPY Resolution:3.80 Å Release Date: 2025-11-19 Classification: MEMBRANE PROTEIN |

