Search Count: 15
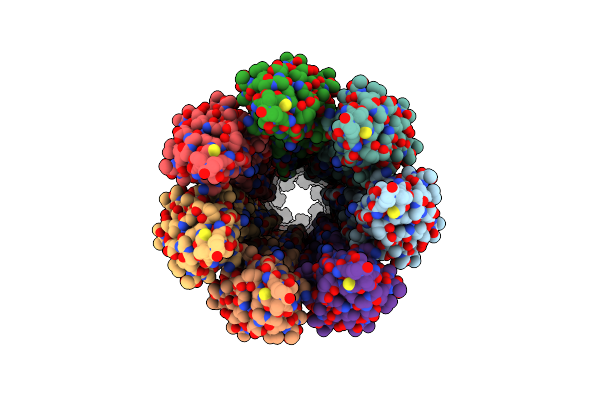 |
Cryoem Structure Of The Neck (1403-2314) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
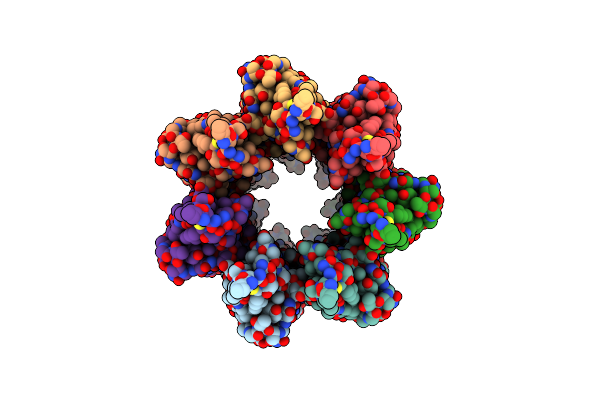 |
Cryoem Structure Of The Fragment-3 (2061-2397) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
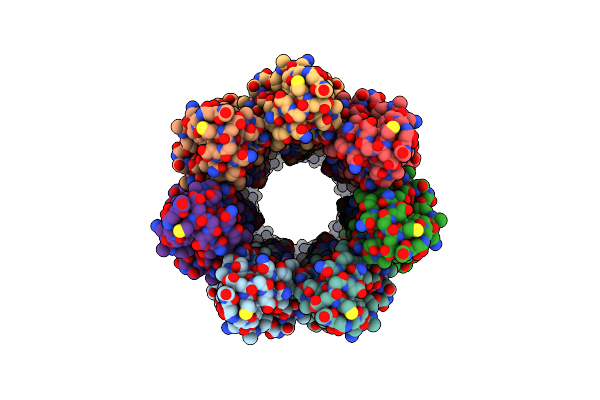 |
Cryoem Structure Of The Fragment-1 (3402-3733) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
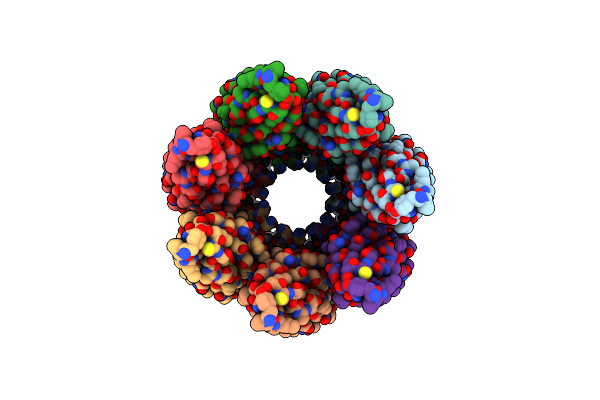 |
Cryoem Structure Of The Fragment-5 (2393-2807) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
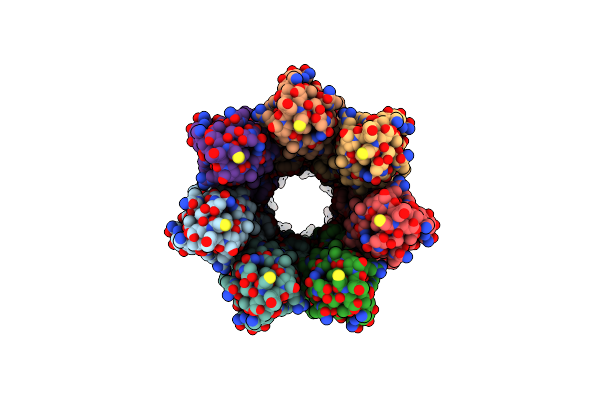 |
Cryoem Structure Of The Fragment-6 (3571-4078) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
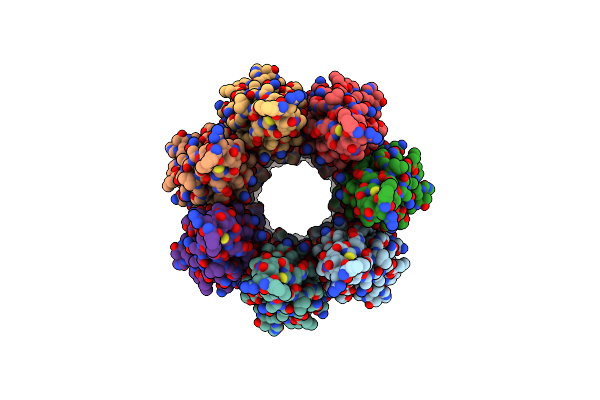 |
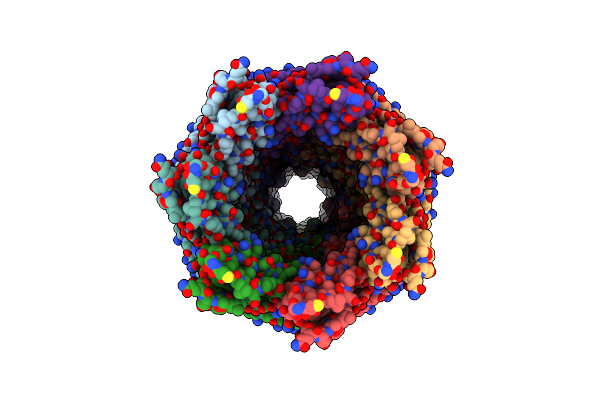
Cryoem Structure Of The Fragment-4 (4074-4421) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
 |
Cryoem Structure Of The Base (4151-5009) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
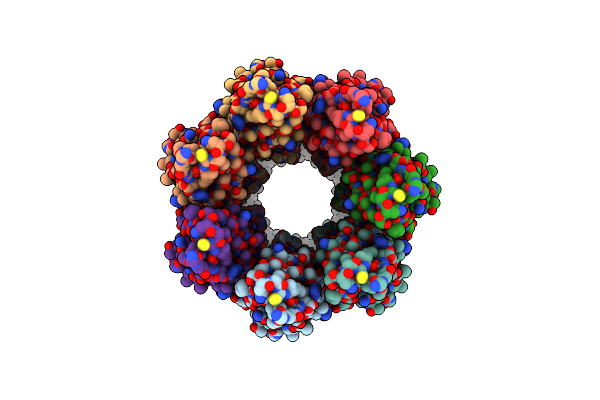 |
Cryoem Structure Of The Fragment-2 (2892-3236) In The Grappling Hook Protein A (Ghpa) In The Bacterium Aureispira Sp. Ccb-Qb1
Organism: Aureispira sp. ccb-qb1
Method: ELECTRON MICROSCOPY Release Date: 2024-10-16 Classification: CELL ADHESION |
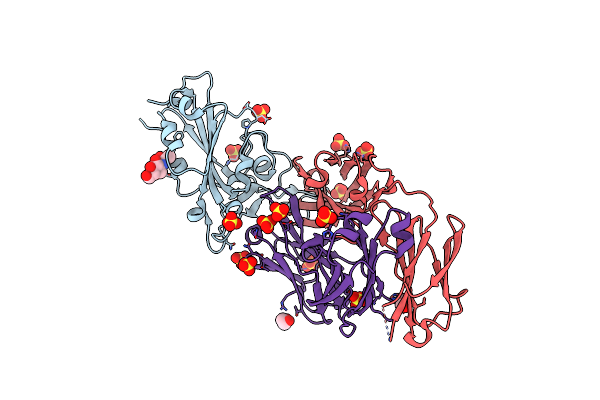 |
Crystal Structure Of Sars-Cov-2 B.1.351 Variant Receptor Binding Domain In Complex With Neutralizing Antibody Cs23
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.86 Å Release Date: 2022-03-09 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG, SO4, EDO |
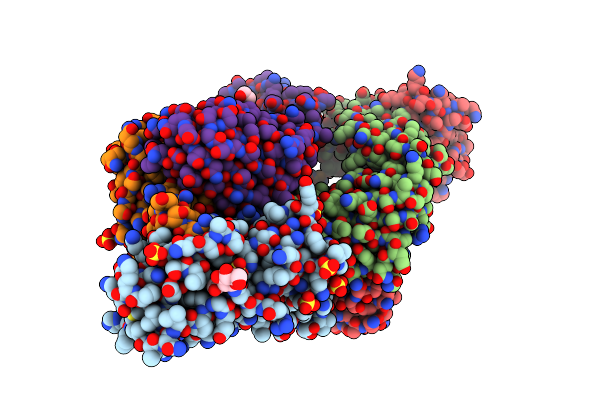 |
Crystal Structure Of Sars-Cov-2 B.1.351 Variant Receptor Binding Domain In Complex With Neutralizing Antibodies Cs44 And Cova1-16
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.88 Å Release Date: 2022-03-09 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG, EDO, SO4 |
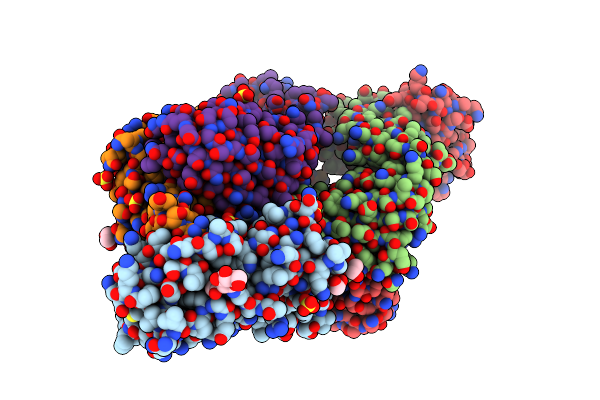 |
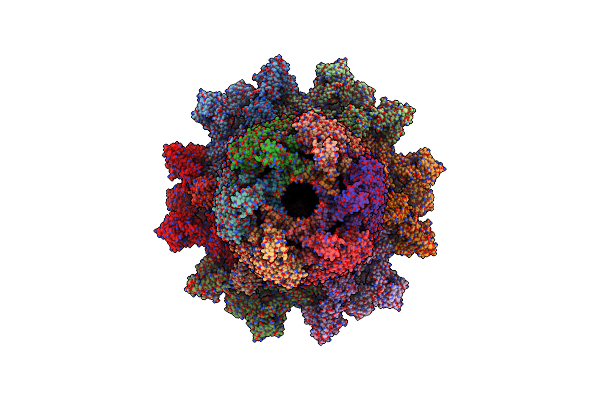
Crystal Structure Of Sars-Cov-2 Receptor Binding Domain In Complex With Neutralizing Antibodies Cv07-287 And Cova1-16
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2022-03-09 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: EDO, NAG, SO4 |
 |
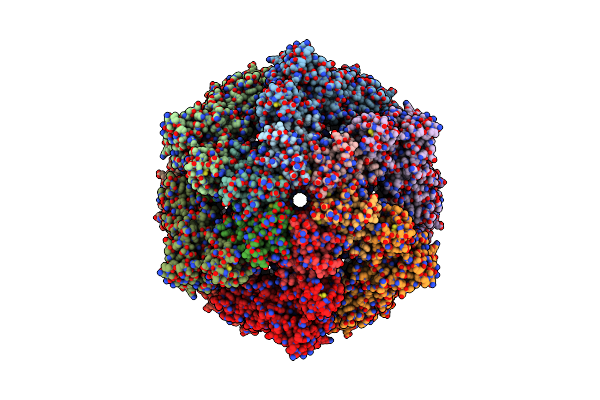
Cryo-Em Structure Of The Contractile Injection System Base Plate From Anabaena Pcc7120
Organism: Nostoc sp. (strain pcc 7120 / sag 25.82 / utex 2576)
Method: ELECTRON MICROSCOPY Release Date: 2022-02-23 Classification: PROTEIN TRANSPORT |
 |
Cryo-Em Structure Of The Contractile Injection System Cap Complex From Anabaena Pcc7120
Organism: Nostoc sp. (strain pcc 7120 / sag 25.82 / utex 2576)
Method: ELECTRON MICROSCOPY Release Date: 2022-02-23 Classification: PROTEIN TRANSPORT |
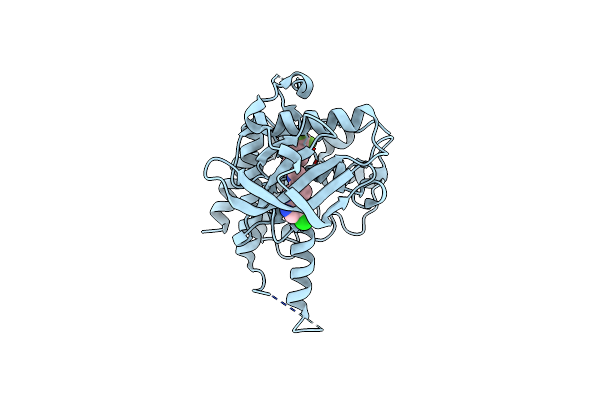 |
Crystal Structure Of Fms Kinase Domain With A Small Molecular Inhibitor, Plx3397
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2015-08-12 Classification: transferase/transferase inhibitor Ligands: P31 |
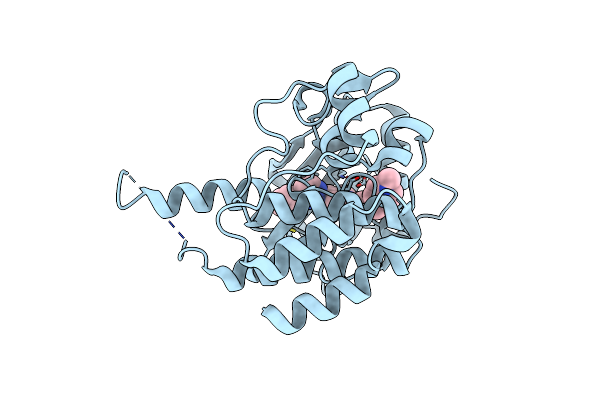 |
Crystal Structure Of Fms Kinase Domain With A Small Molecular Inhibitor, Gleevec
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2015-08-12 Classification: transferase/transferase inhibitor Ligands: STI |

