Search Count: 11
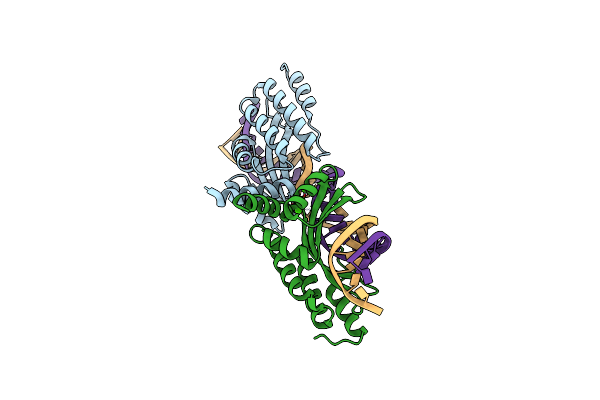 |
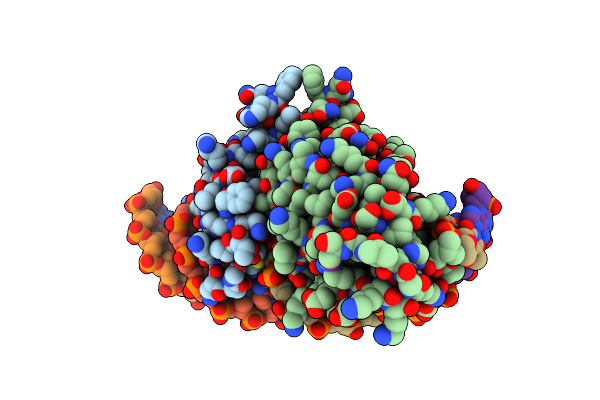
Crystal Structure Of The Homing Endonuclease I-Cvui In Complex With Its Target (Sro1.3) In The Presence Of 2 Mm Ca
Organism: Chlorella vulgaris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2015-09-23 Classification: HYDROLASE/DNA Ligands: CA |
 |
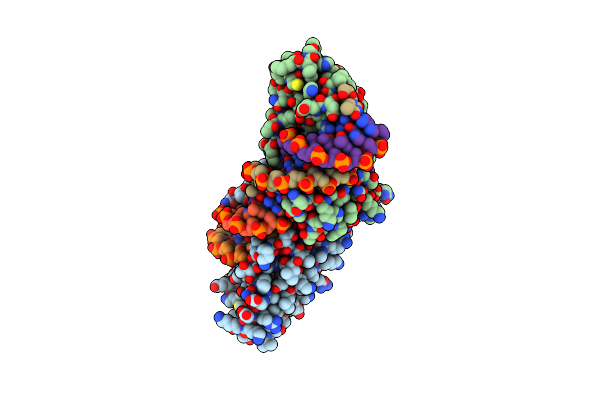
Crystal Structure Of The Homing Endonuclease I-Cvui In Complex With Its Target (Sro1.3) In The Presence Of 2 Mm Mn
Organism: Chlorella vulgaris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2015-09-23 Classification: HYDROLASE/DNA Ligands: MN |
 |
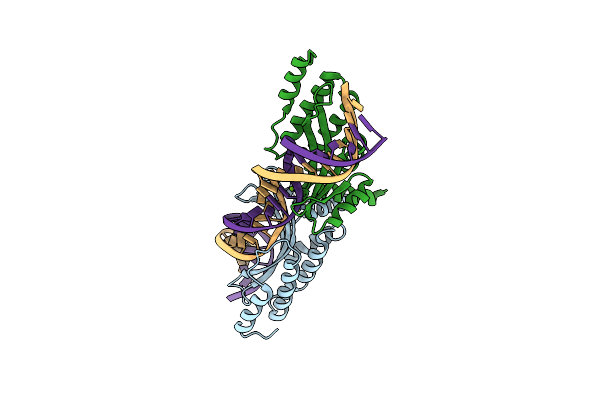
Crystal Structure Of The Homing Endonuclease I-Cvui In Complex With I- Crei Target (C1221) In The Presence Of 2 Mm Mg Revealing Dna Cleaved
Organism: Chlorella vulgaris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2015-09-23 Classification: HYDROLASE/DNA Ligands: MG |
 |
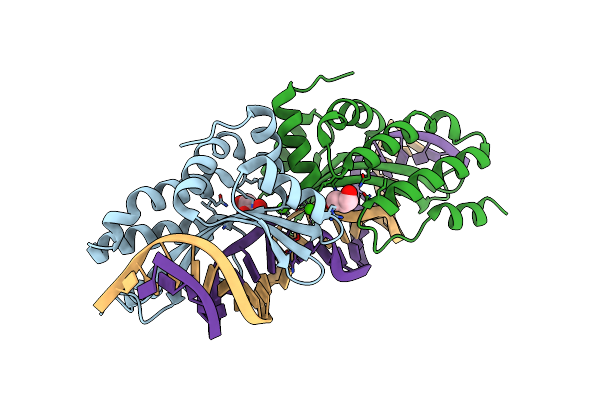
Crystal Structure Of The Homing Endonuclease I-Cvui In Complex With I- Crei Target (C1221) In The Presence Of 2 Mm Mg Revealing Dna Not Cleaved
Organism: Chlorella vulgaris, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2015-09-23 Classification: HYDROLASE/DNA Ligands: MG |
 |
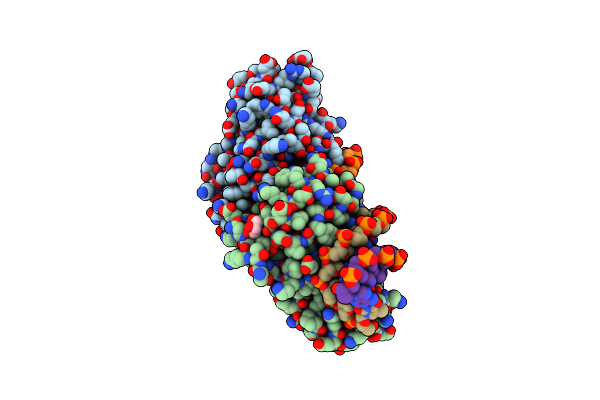
Crystal Structure Of I-Crei Complexed With Its Target Methylated At Position Plus 2 (In The B Strand) In The Presence Of Calcium
Organism: Chlamydomonas reinhardtii, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2012-07-04 Classification: HYDROLASE Ligands: CA, GOL |
 |
Crystal Structure Of I-Crei Complexed With Its Target Methylated At Position Plus 2 (In The B Strand) In The Presence Of Magnesium
Organism: Chlamydomonas reinhardtii, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2012-07-04 Classification: HYDROLASE Ligands: GOL, MG |
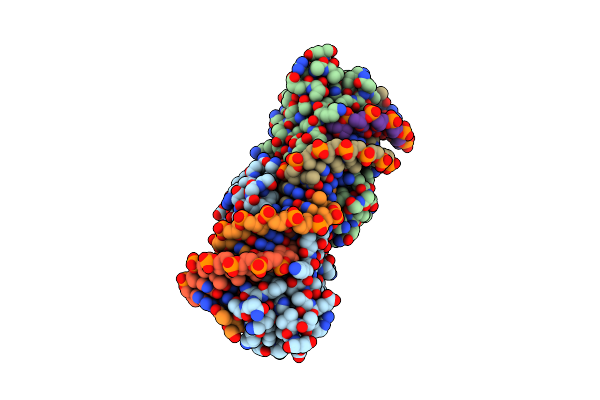 |
Crystal Structure Of The Mutant D75N I-Crei In Complex With Its Wild- Type Target (The Four Central Bases, 2Nn Region, Are Composed By Gtac From 5' To 3')
Organism: Chlamydomonas reinhardtii, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2012-05-02 Classification: HYDROLASE/DNA Ligands: PGO, MG, NA |
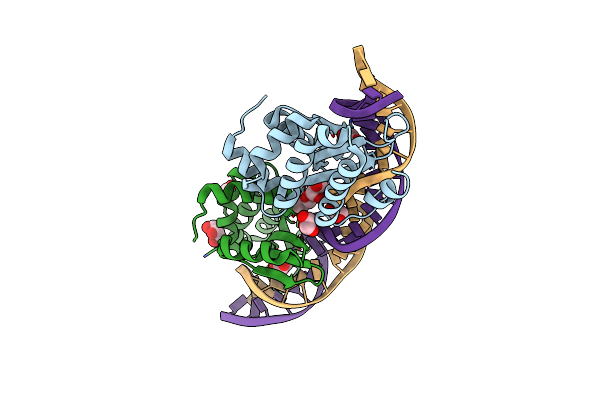 |
Crystal Structure Of The Mutant D75N I-Crei In Complex With Its Wild- Type Target In Absence Of Metal Ions At The Active Site (The Four Central Bases, 2Nn Region, Are Composed By Gtac From 5' To 3')
Organism: Chlamydomonas reinhardtii, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2012-05-02 Classification: HYDROLASE/DNA Ligands: GOL |
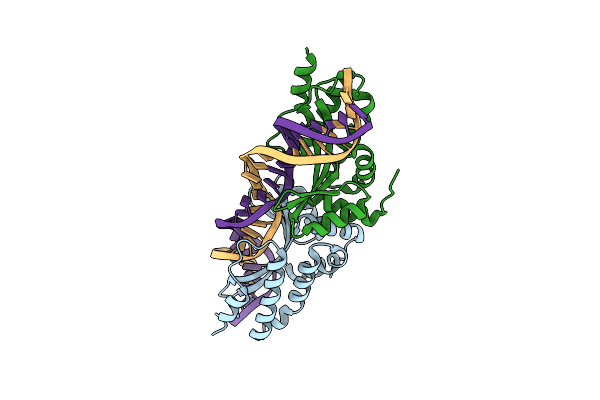 |
Crystal Structure Of The Mutant D75N I-Crei In Complex With An Altered Target (The Four Central Bases, 2Nn Region, Are Composed By Agcg From 5' To 3')
Organism: Chlamydomonas reinhardtii, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2012-05-02 Classification: HYDROLASE/DNA |
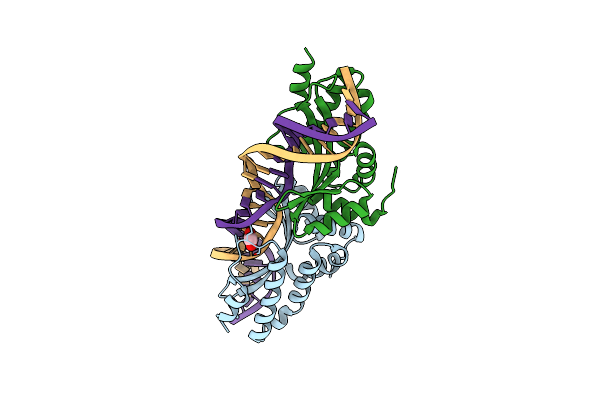 |
Crystal Structure Of The Mutant D75N I-Crei In Complex With An Altered Target (The Four Central Bases, 2Nn Region, Are Composed By Tgca From 5' To 3')
Organism: Chlamydomonas reinhardtii, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2012-05-02 Classification: HYDROLASE/DNA Ligands: EDO |
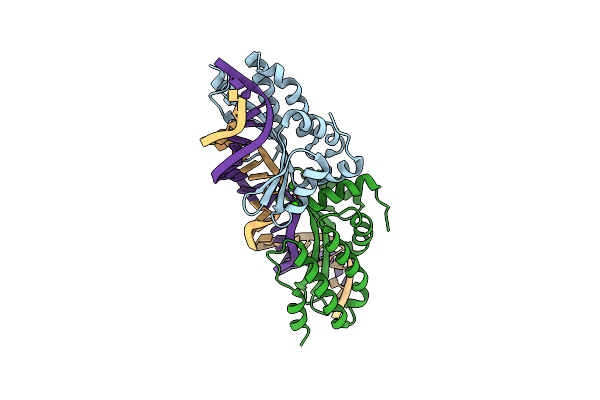 |
Crystal Structure Of The Mutant D75N I-Crei In Complex With Its Wild- Type Target In Presence Of Ca At The Active Site (The Four Central Bases, 2Nn Region, Are Composed By Gtac From 5' To 3')
Organism: Chlamydomonas reinhardtii, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2012-05-02 Classification: HYDROLASE/DNA Ligands: CA |

