Search Count: 775
All
Selected
 |
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: CD, NA, B3P, CIT, EDO, PGE |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) Bound To Neu5Ac2En (Dana)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: DAN, EDO, PEG, CD |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) Bound To Neu5Ac (Nana)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: CD, SLB, EDO, PEG, PGE |
 |
Crystal Structure Of Treponema Denticola Sialidase (Tde_0471) D165A Mutant Bound To 3'-Sialyllactose (Only Neu5Ac Visible)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-02-25 Classification: HYDROLASE Ligands: SIA, GOL, PEG, CD, NA |
 |
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-02-12 Classification: HYDROLASE Ligands: E64, EDO |
 |
Crystal Structure Of C412S Mutant Of C0362 (Tde_0362 [Tde0362] Resi 205-647)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2025-02-12 Classification: HYDROLASE Ligands: EDO, PO4, NA |
 |
Crystal Structure Of Y559A Prosegment Binding Loop Mutant Of C0362 (Tde_0362 [Tde0362] Resi 205-647)
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2025-02-12 Classification: HYDROLASE Ligands: EDO, CL |
 |
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2024-11-27 Classification: HYDROLASE Ligands: K |
 |
Lacto-N-Biosidase From Treponema Denticola Atcc 35405, Histag Bound In Active Site
Organism: Treponema denticola atcc 35405
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2024-10-09 Classification: HYDROLASE Ligands: ZN |
 |
Organism: Treponema denticola atcc 35405
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2024-10-09 Classification: HYDROLASE Ligands: IMD, NA, ZN |
 |
Organism: Treponema denticola atcc 35405
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2024-06-26 Classification: METAL BINDING PROTEIN |
 |
Organism: Treponema denticola atcc 35405
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2024-06-26 Classification: METAL BINDING PROTEIN Ligands: SO4, CU |
 |
Organism: Treponema denticola atcc 35405
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2024-06-26 Classification: METAL BINDING PROTEIN Ligands: SO4, FE, CU |
 |
Organism: Treponema denticola (strain atcc 35405 / dsm 14222 / cip 103919 / jcm 8153 / kctc 15104), Moraxella bovoculi
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-03-22 Classification: GENE REGULATION |
 |
Organism: Treponema denticola (strain atcc 35405 / dsm 14222 / cip 103919 / jcm 8153 / kctc 15104)
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2022-07-13 Classification: HYDROLASE Ligands: MN, CL |
 |
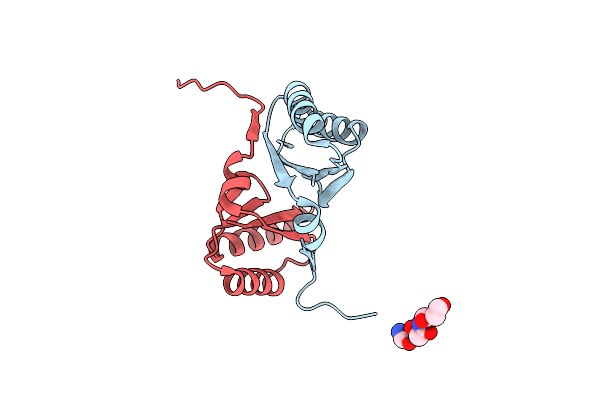
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2020-10-28 Classification: SIGNALING PROTEIN |
 |
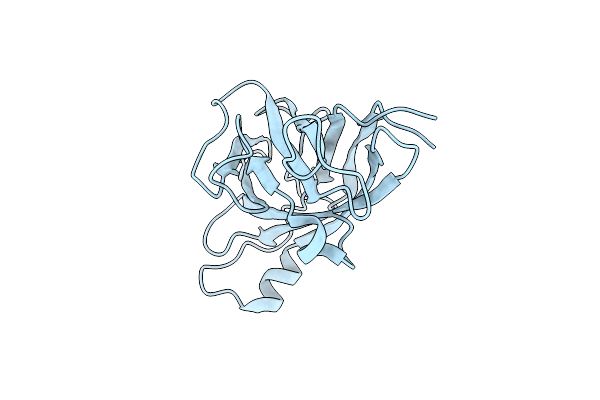
Organism: Treponema denticola (strain atcc 35405 / cip 103919 / dsm 14222), Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2020-03-25 Classification: RNA BINDING PROTEIN |
 |
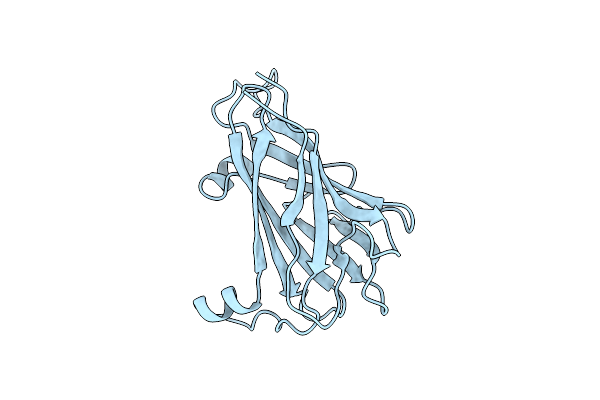
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2019-08-21 Classification: MOTOR PROTEIN |
 |
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2019-08-14 Classification: MOTOR PROTEIN |
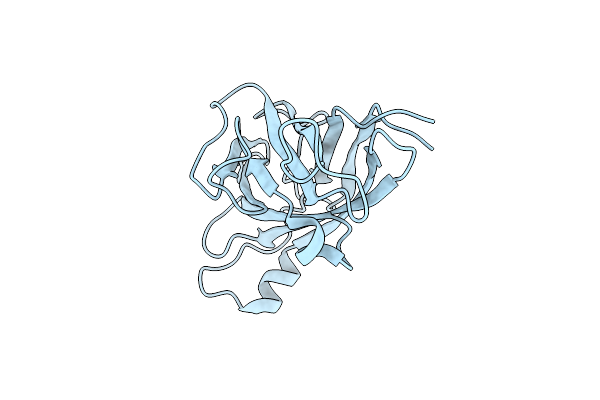 |
Crystal Structure Of The Wild Type D2 Domain (A168-T344) Of The Flagellar Hook Protein Flge From Treponema Denticola
Organism: Treponema denticola
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2019-08-14 Classification: MOTOR PROTEIN |

