Search Count: 5,567
All
Selected
 |
Octamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Nonamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Decamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Cryo-Em Structure Of The Magnesium Transporter Mgta In E1-Like And E2-P Conformations At 2.59 Angstroms
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Release Date: 2026-02-18 Classification: TRANSPORT PROTEIN Ligands: MG, BEF |
 |
Cryo-Em Structure Of The Magnesium Transporter Mgta In The E2 Conformation With Bound Beryllium Fluoride (Bef3-) And Mg2+ At 2.65 A Resolution.
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Resolution:2.59 Å Release Date: 2026-02-18 Classification: TRANSPORT PROTEIN Ligands: MG, BEF |
 |
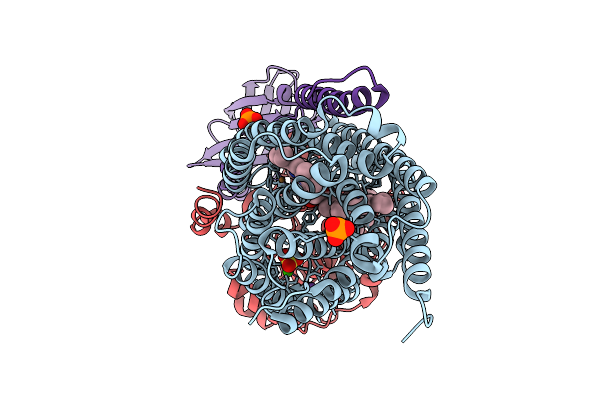
The Cryo-Em Structure Of Hera-Nura Complex With Amppnp From Thermococcus Kodakarensis
Organism: Thermococcus kodakarensis
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-02-04 Classification: TRANSLOCASE Ligands: ANP |
 |
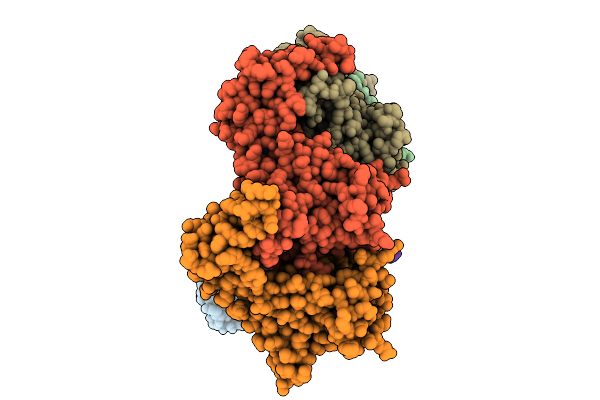
Organism: Stutzerimonas stutzeri atcc 14405 = ccug 16156
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-01-28 Classification: OXIDOREDUCTASE Ligands: HEM, CU, CA, PO4, NA, SO4, HEC |
 |
Pentamer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate State2
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
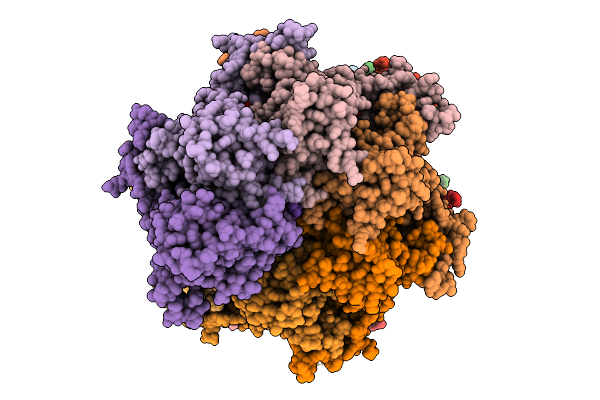
Sixteen Polymer Msp1 From S.Cerevisiae (With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: ATP, MG |
 |
Twenty-Two Polymer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: MG, ATP |
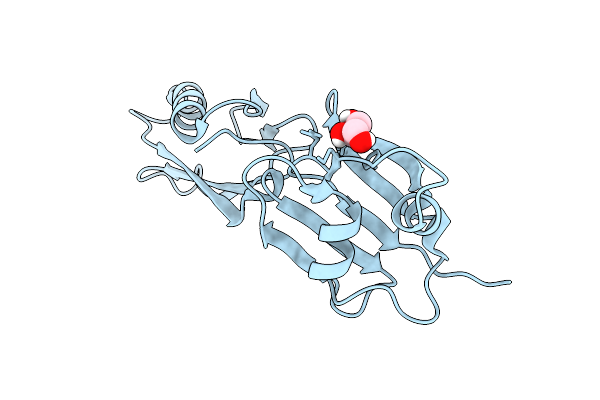 |
Organism: Myxococcus xanthus dz2
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-28 Classification: TRANSLOCASE Ligands: GOL |
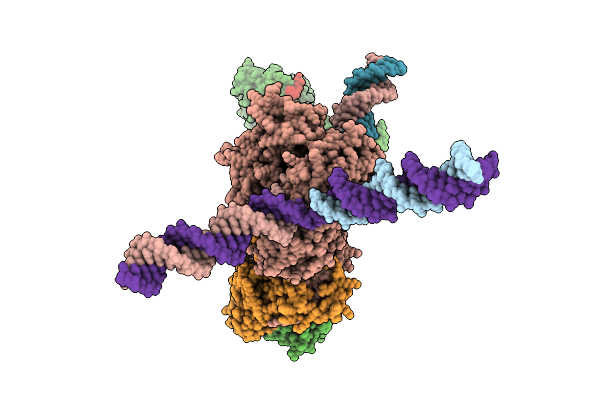 |
Cryo-Em Structure Of F-Box Helicase 1 (Fbh1) Bound To An Scf Ubiquitin Ligase Complex And A 3-Way Dna Fork (Consensus Structure)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: ISOMERASE/DNA Ligands: AGS, MG, ZN |
 |
Cryo-Em Structure Of F-Box Helicase 1 (Fbh1) Bound To An Scf Ubiquitin Ligase Complex And A 3-Way Dna Fork (Head Structure)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: TRANSFERASE |
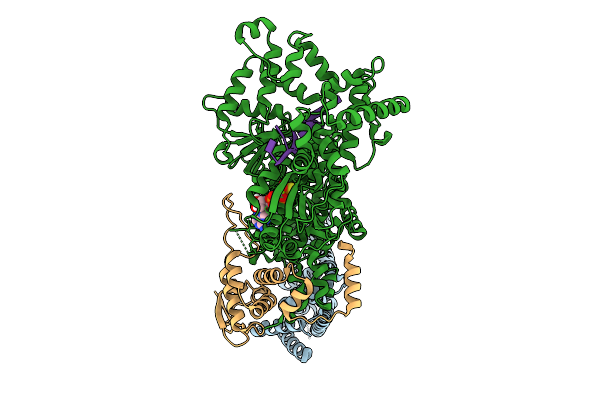 |
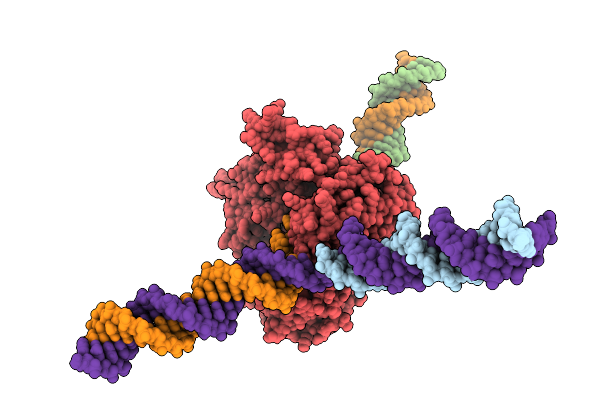
Cryo-Em Structure Of F-Box Helicase 1 (Fbh1) Bound To An Scf Ubiquitin Ligase Complex And A 3-Way Dna Fork (Body Structure)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: ISOMERASE/DNA Ligands: AGS, MG, ZN |
 |
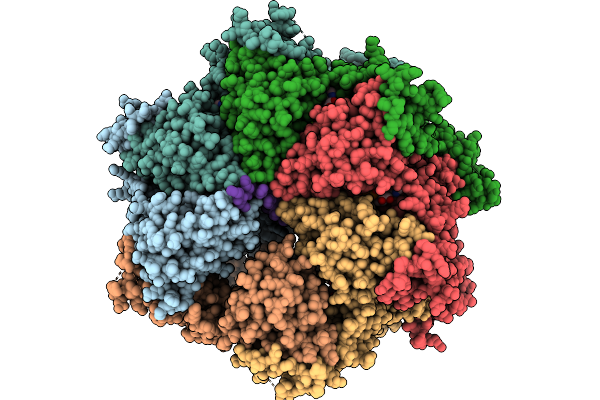
Cryo-Em Structure Of F-Box Helicase 1 (Fbh1) Bound To An Scf Ubiquitin Ligase Complex And A 3-Way Dna Fork (Substrate Structure)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: ISOMERASE/DNA Ligands: AGS, MG, ZN |
 |
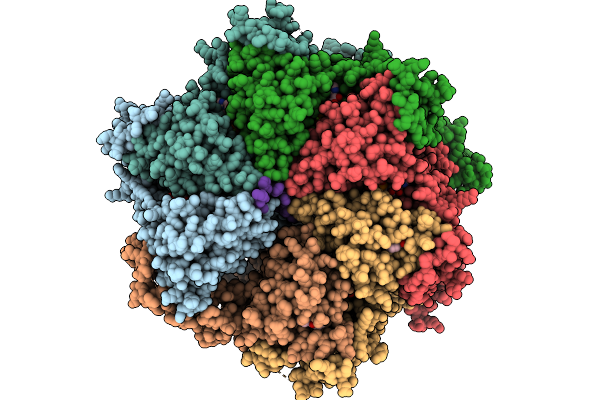
Organism: Escherichia coli bl21(de3), Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: TRANSLOCASE Ligands: ATP, MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: TRANSLOCASE Ligands: ATP, MG, A1AP0 |
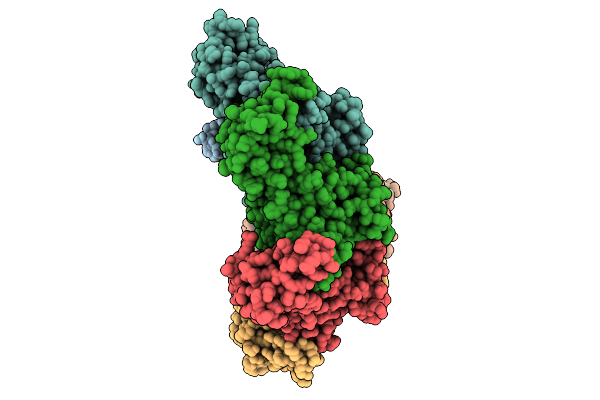 |
Hexamer Msp1 From S.Cerevisiae(With A Catalytic Dead Mutation) In Complex With An Unknown Peptide Substrate
Organism: Saccharomyces cerevisiae s288c, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: MG, ATP |
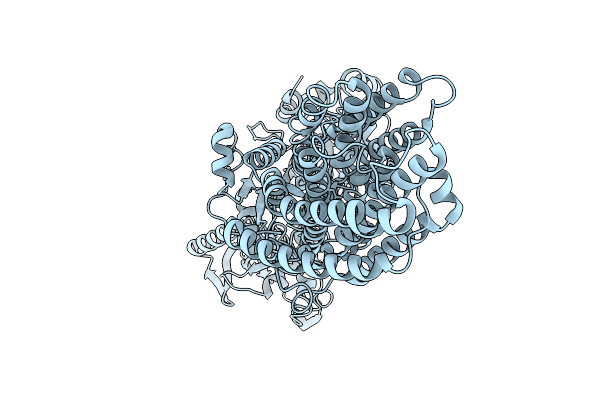 |
Cryo-Em Structure Of The Magnesium Transporter Mgta In The E2 Conformation Bound To Mg2+
Organism: Lactococcus lactis subsp. lactis
Method: ELECTRON MICROSCOPY Resolution:3.23 Å Release Date: 2026-01-21 Classification: TRANSPORT PROTEIN Ligands: MG |
 |

