Search Count: 22
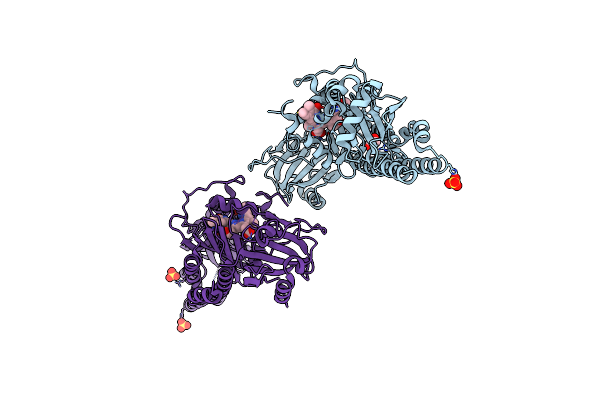 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State (I0A).
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL, EDO |
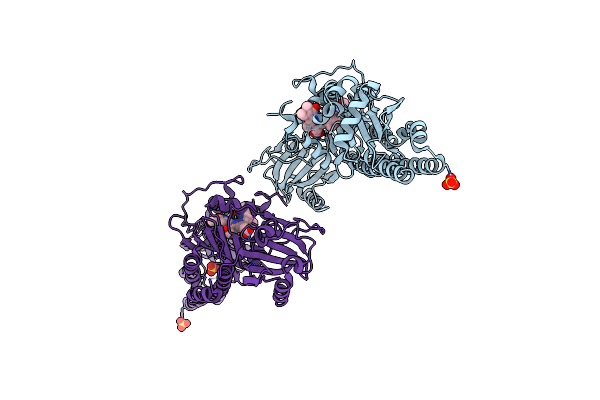 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State (I0B).
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4 |
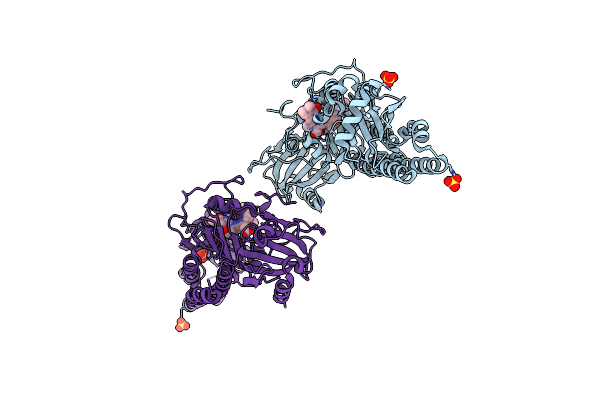 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I1 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4 |
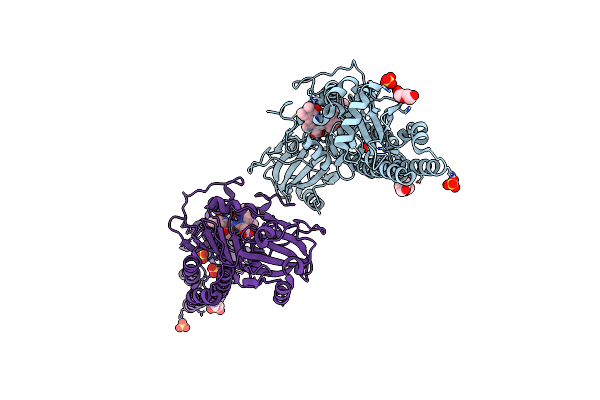 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I2 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, PGE, PEG, CL |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I3 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, GOL, PEG |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I4 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL, PEG |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I5 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I6 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I7 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, PEG |
 |
The Surface-Engineered Photosensory Module (Pas-Gaf-Phy) Of The Bacterial Phytochrome Agp1 (Atbphp1) In The Pr Form With Parallel Dimer Formation
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2024-03-06 Classification: SIGNALING PROTEIN Ligands: V8U, MG |
 |
Cryo-Em Structure Of The Agonist Setmelanotide Bound To The Active Melanocortin-4 Receptor (Mc4R) In Complex With The Heterotrimeric Gs Protein At 2.6 A Resolution.
Organism: Homo sapiens, Rattus norvegicus, Bos taurus, Lama glama, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2021-11-17 Classification: SIGNALING PROTEIN Ligands: CA |
 |
Active Melanocortin-4 Receptor (Mc4R)- Gs Protein Complex Bound To Agonist Ndp-Alpha-Msh At 2.86 A Resolution.
Organism: Homo sapiens, Rattus norvegicus, Mus musculus, Lama glama, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2021-11-17 Classification: SIGNALING PROTEIN Ligands: CA |
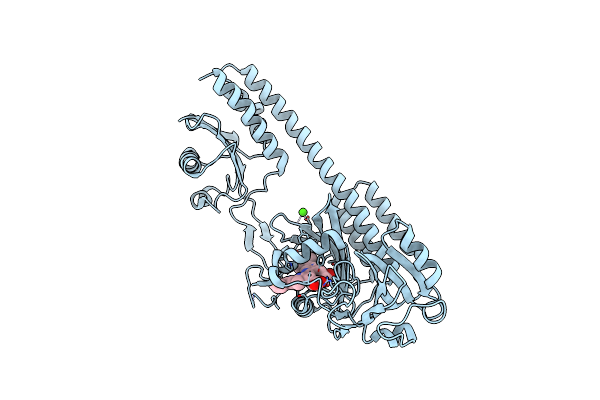 |
The Photosensory Core Module (Pas-Gaf-Phy) Of The Bacterial Phytochrome Agp1 (Atbphp1) Locked In A Pr-Like State
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Release Date: 2020-04-01 Classification: SIGNALING PROTEIN Ligands: JQ2, CA |
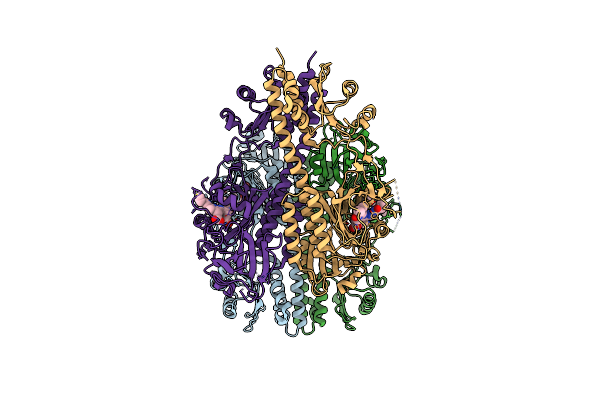 |
Crystallographic Superstructure Of The Photosensory Core Module (Pas-Gaf-Phy) Of The Bacterial Phytochrome Agp1 (Atbphp1) Locked In A Pr-Like State
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Release Date: 2020-04-01 Classification: SIGNALING PROTEIN Ligands: JQ2 |
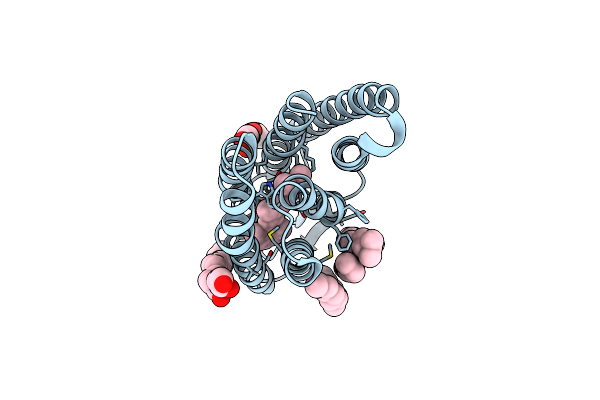 |
Crystal Structure Of The Light-Driven Proton Pump Coccomyxa Subellipsoidea Rhodopsin Csr
Organism: Coccomyxa subellipsoidea c-169
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2019-03-27 Classification: PROTON TRANSPORT Ligands: RET, CLR, OLB |
 |
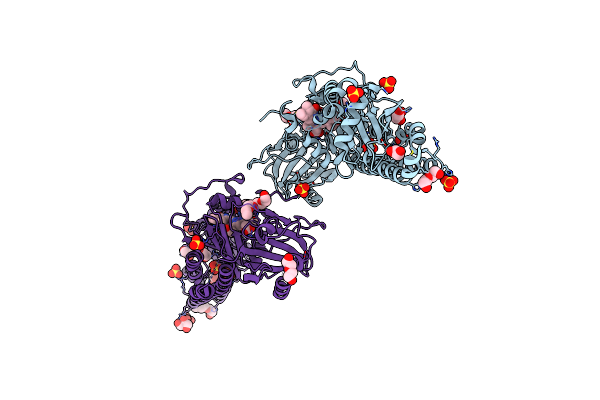
Crystal Structure Of The Photosensory Core Module (Pcm) Of A Bathy Phytochrome From Agrobacterium Fabrum In The Pfr State.
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2018-11-28 Classification: SIGNALING PROTEIN Ligands: EL5 |
 |
Crystal Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State.
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2018-11-28 Classification: FLUORESCENT PROTEIN Ligands: EL5, SO4, GOL, PEG, ETE, PG0 |
 |
Crystal Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Functional Meta-F Intermediate State.
Organism: Agrobacterium fabrum (strain c58 / atcc 33970)
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2018-11-28 Classification: FLUORESCENT PROTEIN Ligands: EL5, SO4, CL, GOL, PEG, ETE, PG0 |
 |
Sfx Structure Of Cydia Pomonella Granulovirus Using A Double Flow-Focusing Nozzle
Organism: Cydia pomonella granulosis virus (isolate mexico/1963)
Method: X-RAY DIFFRACTION Resolution:2.56 Å Release Date: 2017-03-29 Classification: VIRAL PROTEIN |
 |
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: X-RAY DIFFRACTION Resolution:3.80 Å Release Date: 2017-03-29 Classification: TRANSFERASE Ligands: ZN, MG |

