Search Count: 21
 |
Organism: Hypocrea atroviridis (strain atcc 20476 / imi 206040)
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2021-08-04 Classification: LYASE Ligands: 4LU, K, MN |
 |
Organism: Hypocrea atroviridis (strain atcc 20476 / imi 206040)
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2021-08-04 Classification: LYASE Ligands: NO3, BYN, K, MN, ACT |
 |
Structure Of T. Atroviride Variant Tafdcv In Complex With Prfmn-Butynoic Acid Adduct
Organism: Hypocrea atroviridis (strain atcc 20476 / imi 206040)
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2021-08-04 Classification: LYASE Ligands: JRK, K, MN |
 |
Structure Of T. Atroviride Fdc Variant Tafdcv In Complex With Prfmn Crotonic Acid Adduct
Organism: Hypocrea atroviridis (strain atcc 20476 / imi 206040)
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2021-08-04 Classification: LYASE Ligands: ACT, JRH, K, MN |
 |
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2021-08-04 Classification: LYASE Ligands: K, 4LU, MN |
 |
Structure Of A. Niger Fdc T395M R435P P438W Variant (Anfdcii) In Complex With Prfmn
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.69 Å Release Date: 2021-08-04 Classification: LYASE Ligands: MN, K, BYN |
 |
Structure Of The Reversible Pyrrole-2-Carboxylic Acid Decarboxylase Pa0254/Huda
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2021-04-07 Classification: LIGASE Ligands: MN, K, 4LU, IMD |
 |
Structure Of The N318H Variant Of The Reversible Pyrrole-2-Carboxylic Acid Decarboxylase Pa0254/Huda In Complex With Fmn
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2021-04-07 Classification: LIGASE Ligands: FMN, MN, NA |
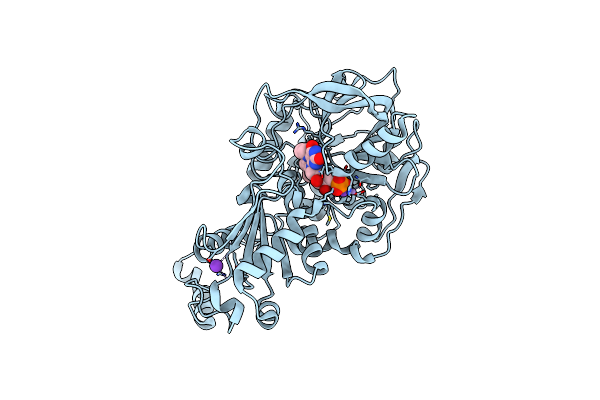 |
Structure Of Wild Type A. Niger Fdc1 Purified In The Dark With Prfmn In The Iminium Form
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.14 Å Release Date: 2017-12-27 Classification: LYASE Ligands: 4LU, MN, K |
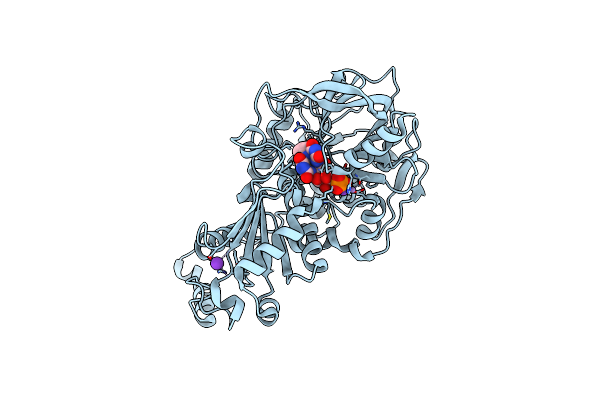 |
Structure Of Wild Type A. Niger Fdc1 That Has Been Illuminated With Uv Light, With Prfmn In The Iminium And Ketimine Form
Organism: Aspergillus niger
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2017-12-20 Classification: LYASE Ligands: 4LU, MN, K, FZZ |
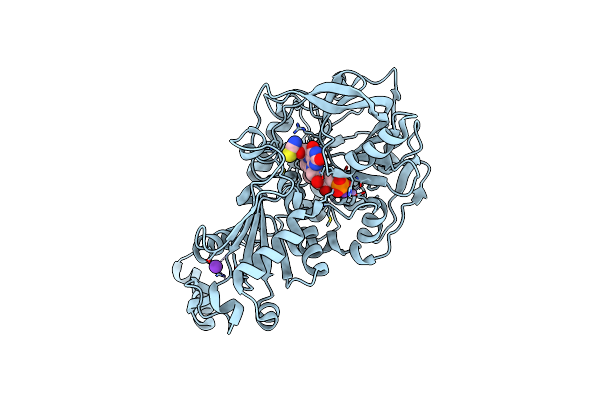 |
Crystal Structure Of E282Q A. Niger Fdc1 With Prfmn In The Hydroxylated Form
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.28 Å Release Date: 2017-12-20 Classification: LYASE Ligands: MN, K, BYN, SCN |
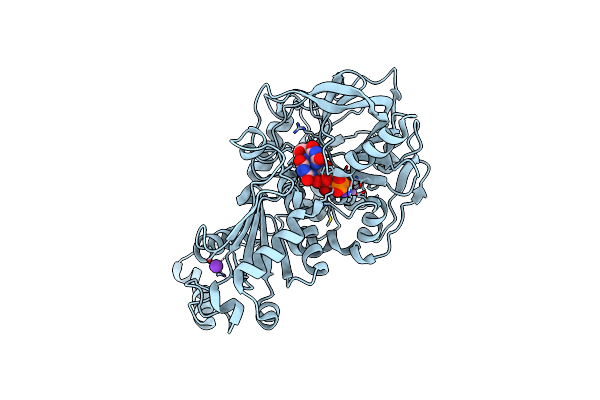 |
Structure Of E282Q A. Niger Fdc1 With Prfmn In The Hydroxylated And Ketimine Forms
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2017-12-20 Classification: LYASE Ligands: MN, K, FZZ, BYN |
 |
Organism: Aspergillus niger
Method: X-RAY DIFFRACTION Resolution:1.06 Å Release Date: 2017-12-20 Classification: LYASE Ligands: MN, K, 4LU |
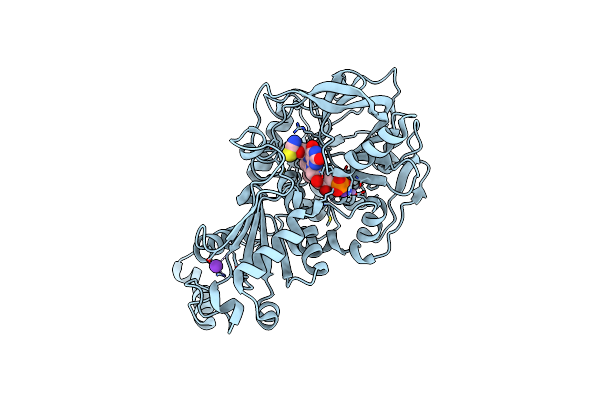 |
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2017-12-20 Classification: LYASE Ligands: MN, K, BYN, SCN |
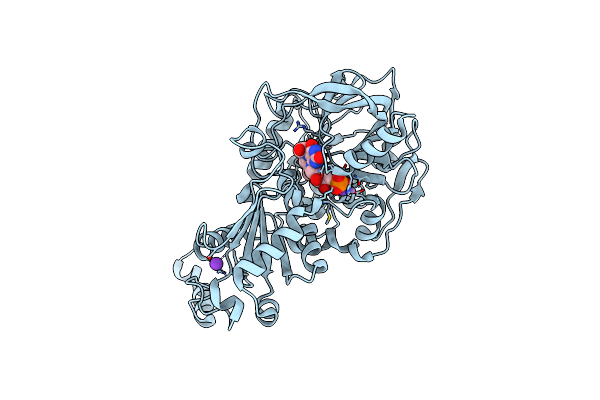 |
Organism: Aspergillus niger
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2017-12-20 Classification: LYASE Ligands: MN, K, BYN |
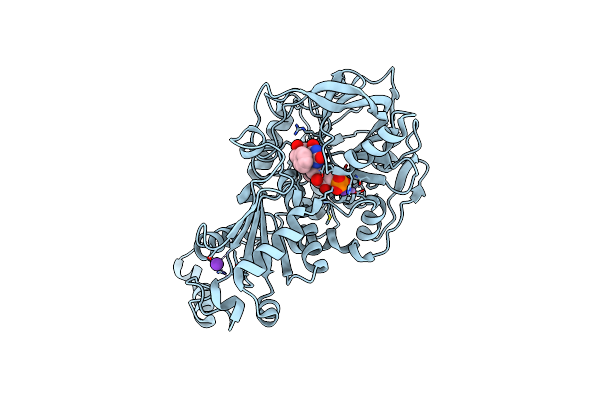 |
Structure Of E277Q A. Niger Fdc1 In Complex With A Phenylpyruvate Derived Adduct To The Prenylated Flavin Cofactor
Organism: Aspergillus niger
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2017-12-20 Classification: LYASE Ligands: MN, K, 4MJ |
 |
Organism: Aspergillus niger
Method: X-RAY DIFFRACTION Resolution:1.13 Å Release Date: 2017-12-20 Classification: LYASE Ligands: 4LU, MN, K, SCN |
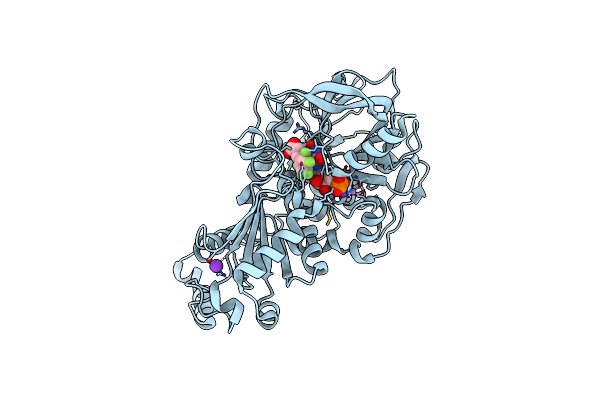 |
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.18 Å Release Date: 2017-12-20 Classification: LYASE Ligands: 4LU, MN, K, F5C |
 |
Organism: Aspergillus niger (strain cbs 513.88 / fgsc a1513)
Method: X-RAY DIFFRACTION Resolution:1.19 Å Release Date: 2017-12-20 Classification: LYASE Ligands: MN, K, BYN |
 |
Organism: Saccharomyces cerevisiae (strain atcc 204508 / s288c)
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2017-12-20 Classification: LYASE Ligands: 4LU, MN, K |

