Search Count: 527
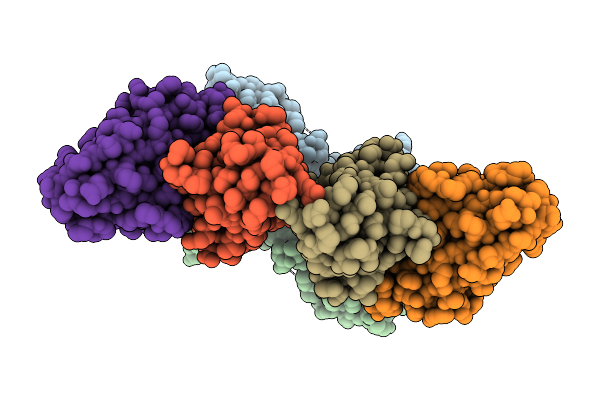 |
Cryo-Em Structure Of Human Complement C1S Cub Domain In Complex With Ray121
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-11-19 Classification: IMMUNE SYSTEM Ligands: CA |
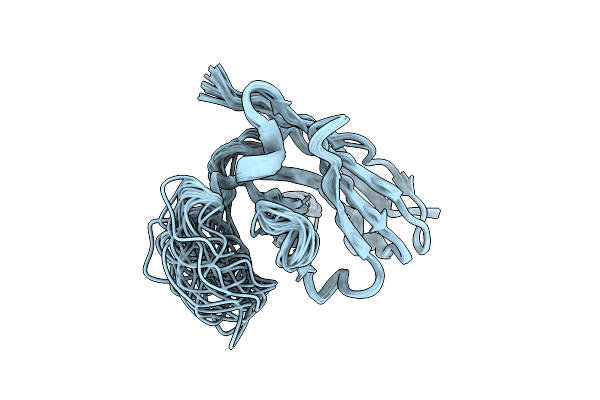 |
Organism: Vicugna pacos
Method: SOLUTION NMR Release Date: 2025-11-12 Classification: PROTEIN BINDING |
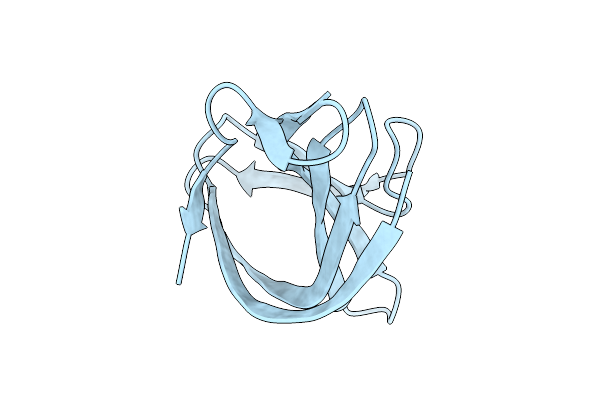 |
Organism: Streptococcus phage philp081102
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: LIGASE |
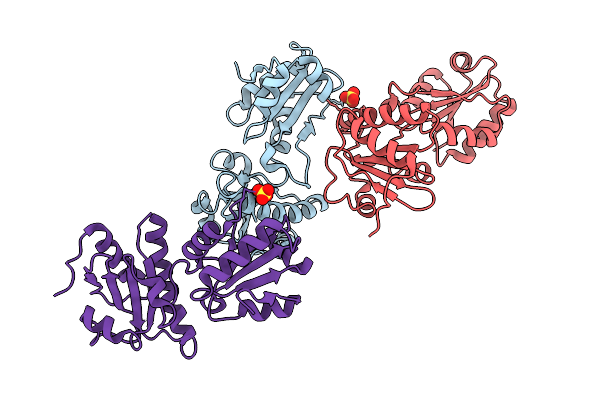 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-10-01 Classification: ISOMERASE Ligands: SO4 |
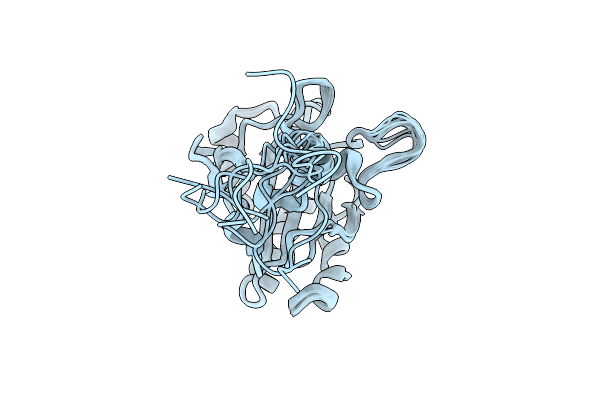 |
Solution Structure Of Human Glutathione Peroxidase 4 (Sec73Cys) With Eight Mutations
|
 |
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2025-08-13 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-07-23 Classification: TRANSFERASE Ligands: A1L3B, CL, GOL, EDO |
 |
 |
Organism: Staphylococcus aureus
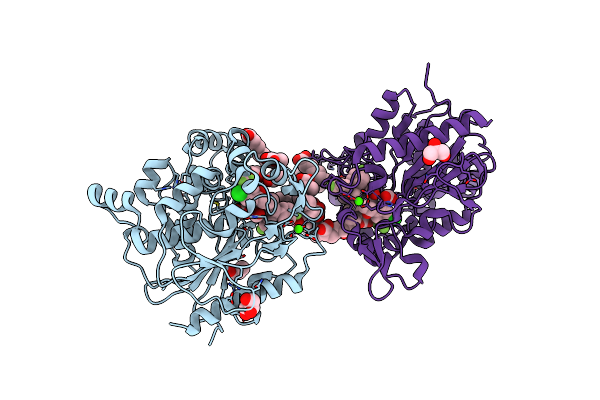
Method: X-RAY DIFFRACTION Release Date: 2025-04-30 Classification: HYDROLASE Ligands: OCA, PPI, GOL, A1L60, FMT, ZN, CA, 6NA, CL, BUA, 11A |
 |
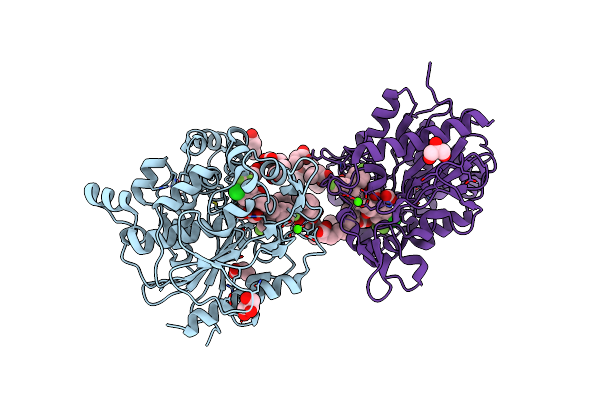
Crystal Structure Of Tyrosine Phenol-Lyase In Complex With 3,5-Dihydroxybenzoic Acid
Organism: Morganella morganii subsp. morganii
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2025-04-23 Classification: LYASE Ligands: 34D, K |
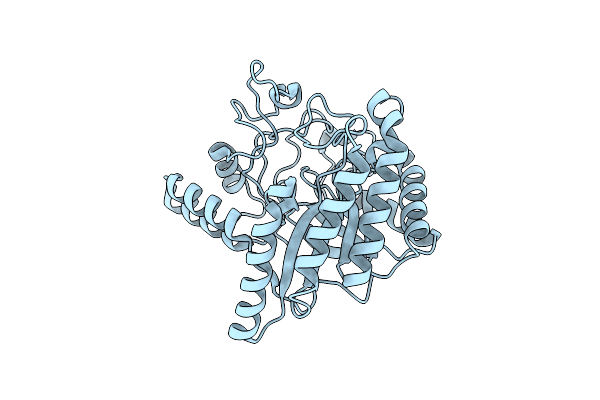 |
Neutron Structure Of Cellulase Cel6A From Phanerochaete Chrysosporium At Room Temperature
Organism: Phanerodontia chrysosporium
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:1.36 Å, 1.86 Å Release Date: 2025-03-12 Classification: HYDROLASE |
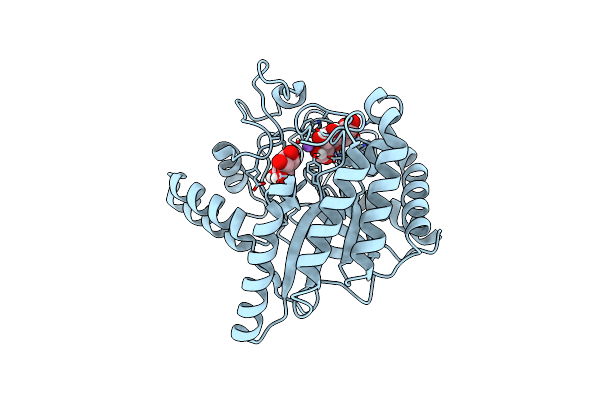 |
Neutron Structure Of Cellulase Cel6A From Phanerochaete Chrysosporium At Room Temperature, Enzyme-Product Complex
Organism: Phanerodontia chrysosporium
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:1.40 Å, 1.8 Å Release Date: 2025-03-12 Classification: HYDROLASE Ligands: BGC, NA |
 |
Neutron Structure Of Cellulase Cel6A From Phanerochaete Chrysosporium At Room Temperature, Low-D2O-Solvent
Organism: Phanerodontia chrysosporium
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:1.40 Å, 2.15 Å Release Date: 2025-03-12 Classification: HYDROLASE |
 |
Neutron Structure Of Cellulase Cel6A From Phanerochaete Chrysosporium At Room Temperature, Enzyme-Product Complex, H2O Solvent
Organism: Phanerodontia chrysosporium
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:1.80 Å, 2.59 Å Release Date: 2025-03-12 Classification: HYDROLASE Ligands: BGC, NA |
 |
High-Resolution X-Ray Structure Of Cellulase Cel6A From Phanerochaete Chrysosporium At Cryogenic Temperature
Organism: Phanerodontia chrysosporium
Method: X-RAY DIFFRACTION Resolution:0.80 Å Release Date: 2025-03-12 Classification: HYDROLASE Ligands: MPD, ACT |
 |
High-Resolution X-Ray Structure Of Cellulase Cel6A From Phanerochaete Chrysosporium At Cryogenic Temperature, Enzyme-Product Complex
Organism: Phanerodontia chrysosporium
Method: X-RAY DIFFRACTION Resolution:0.85 Å Release Date: 2025-03-12 Classification: HYDROLASE Ligands: BGC, NA, CL, PEG |
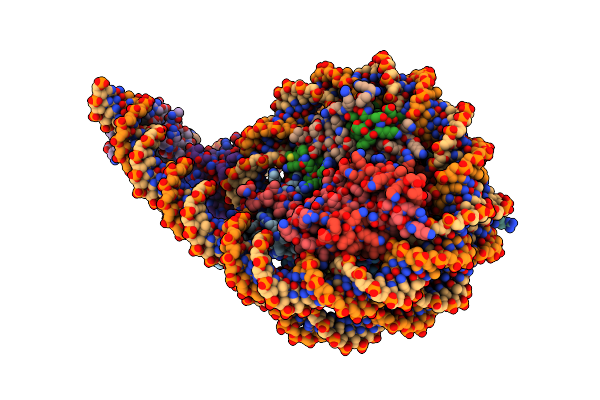 |
Cryo-Em Structure Of Baf-Lamin A/C Igf-Nucleosome Complex (High Mobility Complex)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: NUCLEAR PROTEIN |
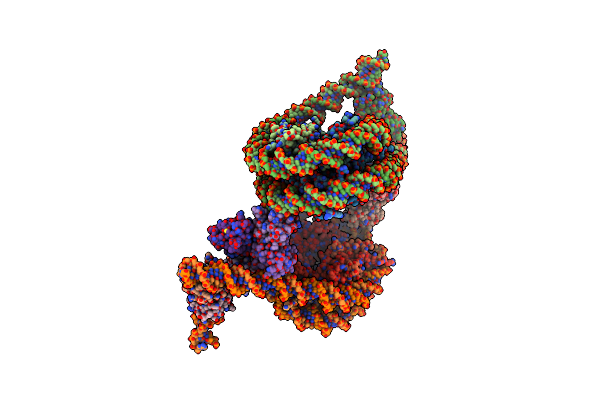 |
Cryo-Em Structure Of Baf-Lamin A/C Igf-Nucleosome Complex (Low Mobility Complex)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: NUCLEAR PROTEIN |
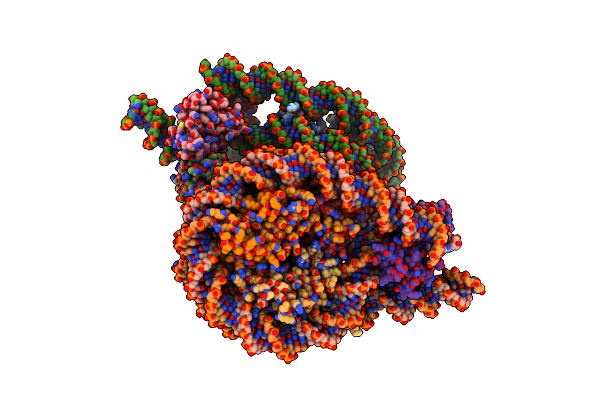 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: NUCLEAR PROTEIN |
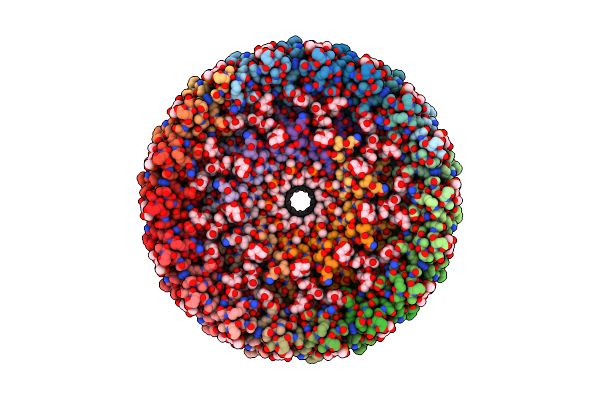 |
Photosynthetic Lh2-Lh1 Complex From The Purple Bacterium Halorhodospira Halophila
Organism: Halorhodospira halophila
Method: ELECTRON MICROSCOPY Release Date: 2025-02-19 Classification: PHOTOSYNTHESIS Ligands: BCL, LMT, CRT, PGV |

