Search Count: 100
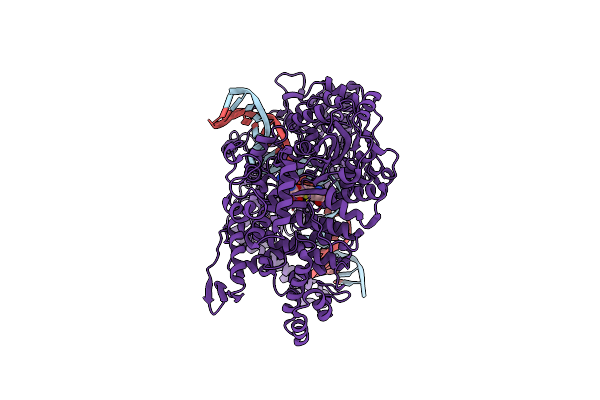 |
Cryo-Em Structure Of The E. Coli Brxx Methyltransferase In Complex With Dna
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: TRANSFERASE Ligands: MG, SAM |
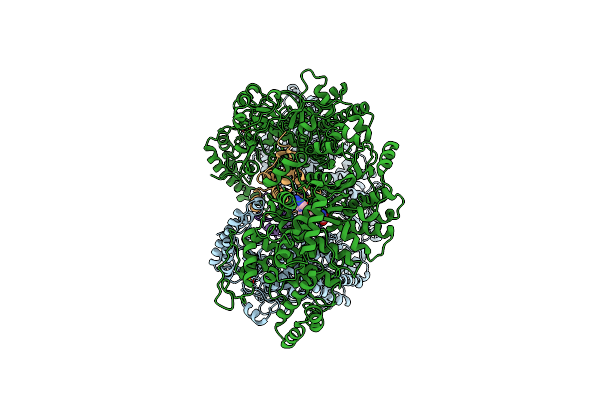 |
Cryo-Em Structure Of The E. Coli Brxx Methyltransferase In Complex With Ocr
Organism: Escherichia coli, Escherichia phage t7
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: TRANSFERASE Ligands: MG, SAM |
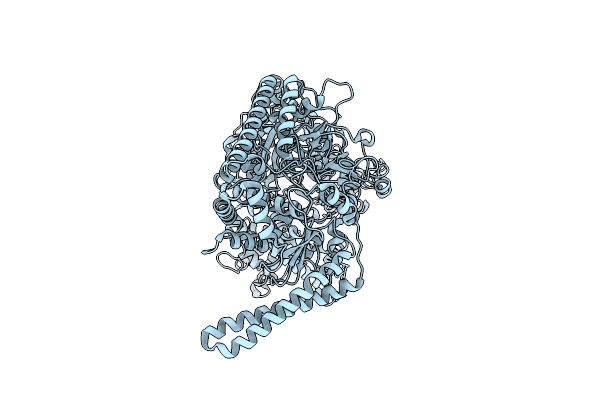 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2025-03-05 Classification: TRANSFERASE |
 |
Organism: Campylobacter jejuni
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-02-12 Classification: TOXIN/ANTITOXIN Ligands: SO4 |
 |
Organism: Escherichia coli b185
Method: ELECTRON MICROSCOPY Release Date: 2024-09-18 Classification: IMMUNE SYSTEM Ligands: AGS |
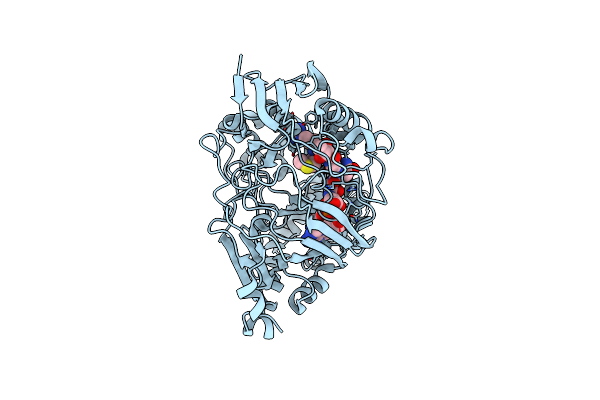 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2024-04-17 Classification: PEPTIDE BINDING PROTEIN/ANTIBIOTIC |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2024-02-07 Classification: TRANSPORT PROTEIN/ANTIBIOTIC |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-02-07 Classification: PEPTIDE BINDING PROTEIN/ANTIBIOTIC |
 |
Organism: Thermus virus p74-26
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2024-01-17 Classification: REPLICATION Ligands: MG |
 |
Organism: Oshimavirus p2345
Method: X-RAY DIFFRACTION Resolution:3.08 Å Release Date: 2023-10-11 Classification: TRANSCRIPTION |
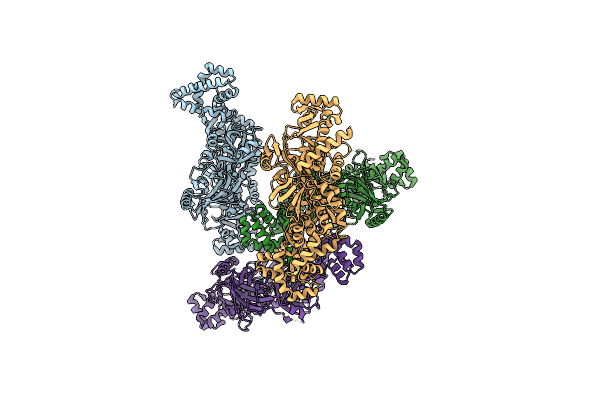 |
Organism: Thermus virus p23-45
Method: X-RAY DIFFRACTION Resolution:4.41 Å Release Date: 2023-10-11 Classification: TRANSCRIPTION Ligands: MG |
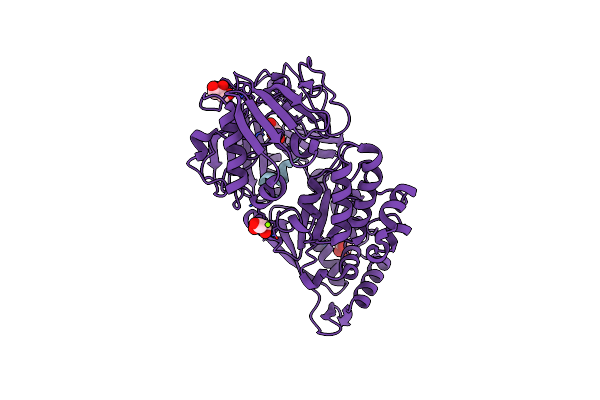 |
Crystal Structure Of The Substrate-Binding Protein Yeja In Complex With Peptide Fragment
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-09-20 Classification: TRANSPORT PROTEIN Ligands: GOL, CL, MG |
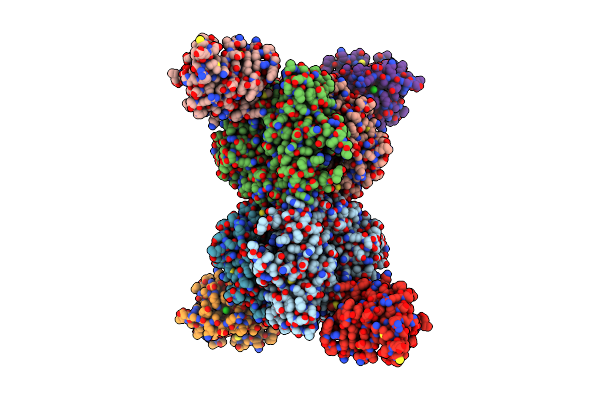 |
Organism: Enterobacteria phage t3, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2023-08-23 Classification: LYASE Ligands: CL |
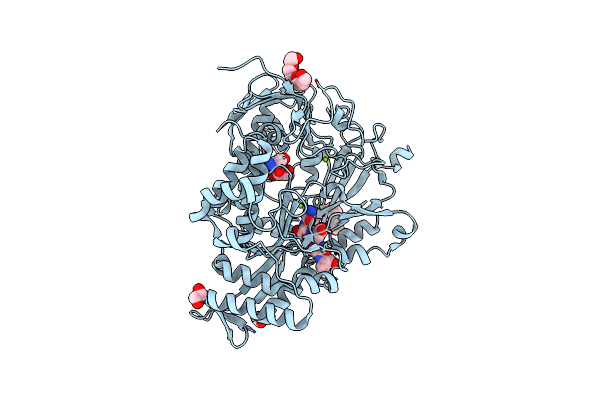 |
Crystal Structure Of The Substrate-Binding Protein Yeja From S. Meliloti In Complex With Peptide Fragment
Organism: Sinorhizobium meliloti
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2023-02-22 Classification: TRANSPORT PROTEIN Ligands: PG4, EDO, MG |
 |
Organism: Bacillus phage ar9
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2022-07-06 Classification: TRANSCRIPTION Ligands: ZN |
 |
X-Ray Structure Of The Phage Ar9 Non-Virion Rna Polymerase Holoenzyme In Complex With A Forked Oligonucleotide Containing The P077 Promoter
Organism: Bacillus phage ar9
Method: X-RAY DIFFRACTION Resolution:3.40 Å Release Date: 2022-07-06 Classification: TRANSCRIPTION/DNA Ligands: ZN |
 |
Structure Of The Phage Ar9 Non-Virion Rna Polymerase Holoenzyme In Complex With Two Dna Oligonucleotides Containing The Ar9 P077 Promoter As Determined By Cryo-Em
Organism: Bacillus phage ar9, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2022-07-06 Classification: TRANSCRIPTION Ligands: ZN |
 |
Structure Of Bacteriophage Ar9 Non-Virion Rnap Polymerase Holoenzyme Determined By Cryo-Em
Organism: Bacillus phage ar9
Method: ELECTRON MICROSCOPY Release Date: 2022-07-06 Classification: TRANSCRIPTION Ligands: ZN |
 |
Organism: Cellulophaga phage phi14:2
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2020-07-29 Classification: VIRAL PROTEIN, TRANSFERASE, TRANSCRIPTION Ligands: CL, NA |
 |
Organism: Pseudomonas phage luz7
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2019-10-30 Classification: VIRAL PROTEIN Ligands: PO4 |

