Search Count: 7,366
All
Selected
 |
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Ala Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.29 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Glu Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.52 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps) In Complex With Rna Microhelices Mimicking Trna-Glu Acceptor Arm
Organism: Nocardia brasiliensis atcc 700358, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:3.43 Å Release Date: 2026-03-04 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
Mycoplasma Penetrans Methionyl Trna Synthetase Is An Asymmetric Dimer Fused To N-Terminal Ancillary Domains
Organism: Malacoplasma penetrans
Method: ELECTRON MICROSCOPY Release Date: 2026-02-25 Classification: LIGASE |
 |
Structure Of Cyclodipeptide Synthase From Nocardia Brasiliensis (Nbra-Cdps)
Organism: Nocardia brasiliensis atcc 700358
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-02-18 Classification: RNA BINDING PROTEIN Ligands: PO4 |
 |
Crystal Structure Of Chimeric Air Synthetase Constructed From E. Coli (Residues 1-168) And Pyrococcus Abyssi (Residues 163-334)
Organism: Escherichia coli, Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: LIGASE |
 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 43-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE Ligands: SO4 |
 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 54-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE |
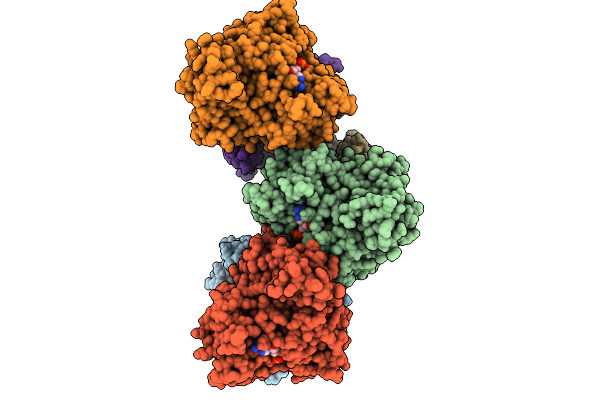 |
Crystal Structure Of Gamma-Glutamyl-Methylamide Synthetase From Methylovorus Mays (Mmgmas) In Complex With Atpgs
Organism: Methylovorus mays
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2026-01-28 Classification: LIGASE Ligands: AGS, MG |
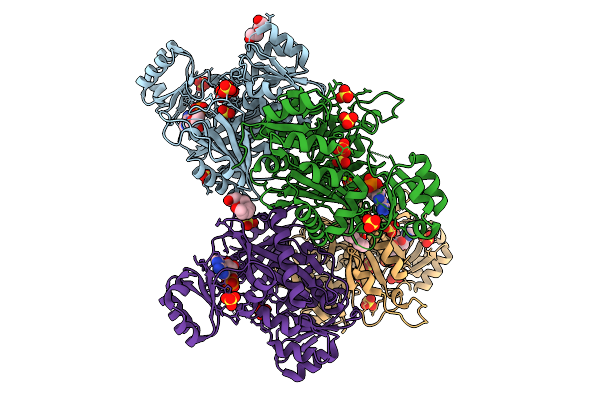 |
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Amp, Adp And Sulfate Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2026-01-21 Classification: LYASE Ligands: AMP, PEG, MG, SO4, PG4, ADP |
 |
The Structure Of Hitb-Hitd Complex With A C4 Pantetheine Cross-Linking Probe
Organism: Embleya scabrispora
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-01-21 Classification: LIGASE Ligands: ADP, GOL, A1L7C |
 |
Organism: Penicillium aurantiogriseum
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-01-14 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL, TRS, MG, ATP |
 |
Organism: Penicillium aurantiogriseum
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-01-14 Classification: BIOSYNTHETIC PROTEIN Ligands: GOL |
 |
Organism: Pseudomonas aeruginosa pao1
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: LIGASE |
 |
Organism: Pseudomonas aeruginosa pao1
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: LIGASE |
 |
Organism: Pseudomonas aeruginosa pao1
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: LIGASE |
 |
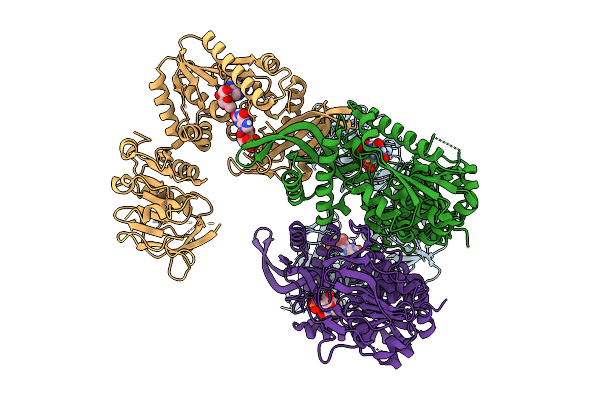
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.93 Å Release Date: 2026-01-07 Classification: LIGASE Ligands: ZN |
 |
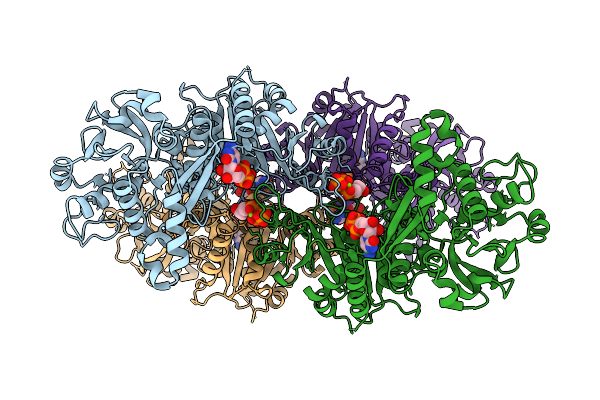
Organism: Escherichia coli k-12
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2025-12-24 Classification: LIGASE/ANTIBIOTIC Ligands: XMP, A1L67 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: STRUCTURAL PROTEIN Ligands: CTP, MG |
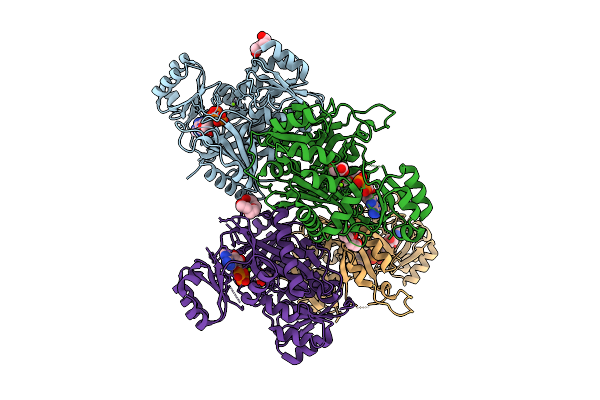 |
Crystal Structure Of Phosphoribosylaminoimidazole Carboxylase From Burkholderia Xenovorans (Atp Complex)
Organism: Paraburkholderia xenovorans lb400
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2025-12-24 Classification: LYASE Ligands: ATP, PEG, MG, PGE, GOL, ADP, PG4 |

