Search Count: 8,493
All
Selected
 |
Organism: Streptomyces phage phi-c31, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-03-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
Organism: Streptomyces xinghaiensis s187
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-04 Classification: BIOSYNTHETIC PROTEIN Ligands: MG, GOL, EDO |
 |
Xylose Isomerase Collected At 20C Using Time-Resolved Serial Synchrotron Crystallography With Glucose At 180 Seconds
Organism: Streptomyces rubiginosus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-02-18 Classification: ISOMERASE Ligands: GLC, GLO, MG |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD, CL |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis (Alternative Loop Conformation)
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis (Alternative N-Terminus)
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Crystal Structure Of Ortho-Aminophenol Oxidase Smnspf From Streptomyces Murayamaensis
Organism: Streptomyces murayamaensis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: CU, CL |
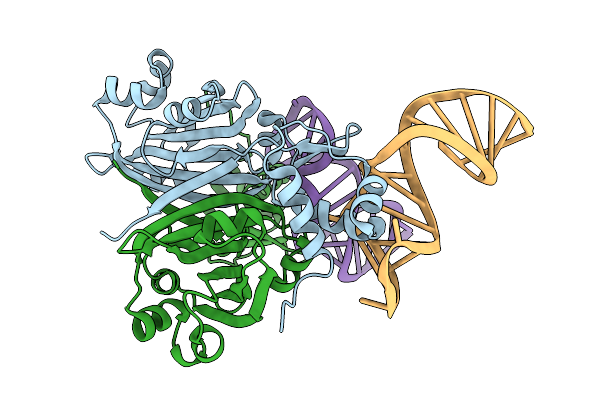 |
Crystal Structure Of Dasr In Complex With A Synthetic Dasr-Binding Rna Aptamer
Organism: Streptomyces coelicolor, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-01-28 Classification: RNA BINDING PROTEIN Ligands: BR |
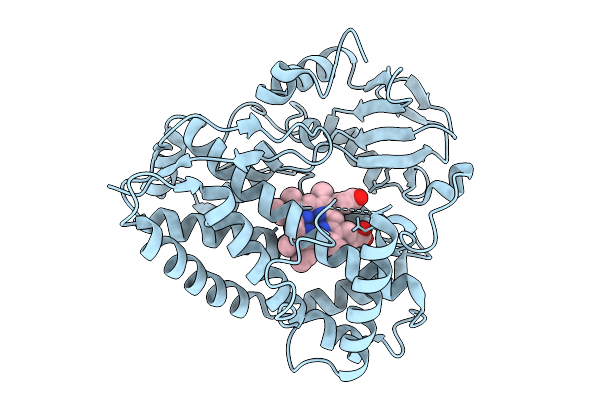 |
Organism: Streptomyces atratus
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-01-28 Classification: METAL BINDING PROTEIN Ligands: HEM |
 |
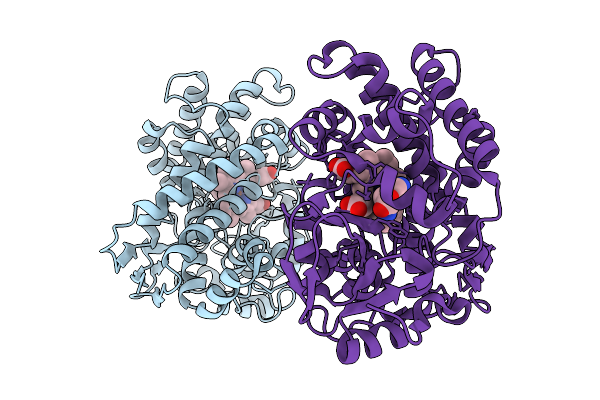
Sam-Dependent C6-Fpp Methytransferase From Streptomyces Varsoviensis In Complex With Sah And Fpp
Organism: Streptomyces varsoviensis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: FPP, SAH, PG4, PGE, PE8, PEG, EDO |
 |
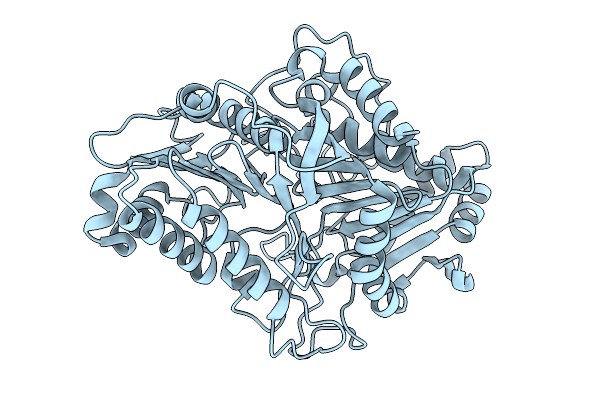
Organism: Streptomyces scabiei 87.22
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2026-01-28 Classification: BIOSYNTHETIC PROTEIN Ligands: TRP, HEM |
 |
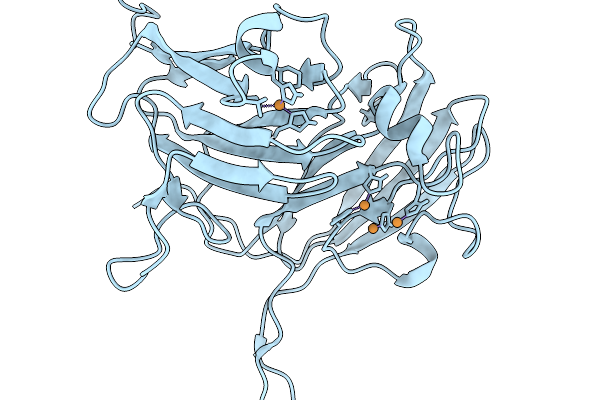
Organism: Streptomyces klenkii
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-01-21 Classification: HYDROLASE |
 |
Organism: Streptomyces coelicolor
Method: X-RAY DIFFRACTION Resolution:3.51 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: CU |
 |
Comparative Analysis Of Functions And Catalytic Mechanisms Of Methyltransferases Involved In Anthracycline Biosynthesis
Organism: Streptomyces coeruleorubidus
Method: X-RAY DIFFRACTION Resolution:1.56 Å Release Date: 2026-01-07 Classification: TRANSFERASE Ligands: SAH, A1L3V |
 |
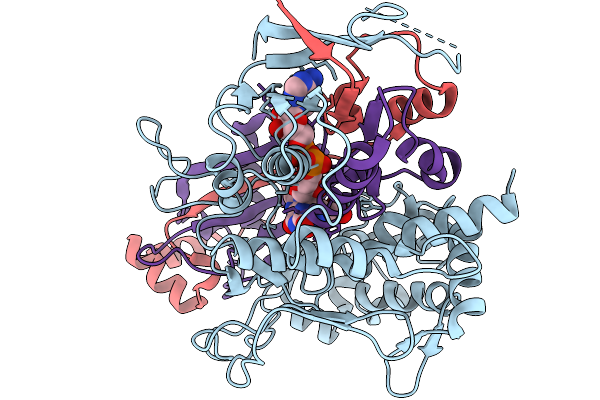
Comparative Analysis Of Functions And Catalytic Mechanisms Of Methyltransferases Involved In Anthracycline Biosynthesis
Organism: Streptomyces coeruleorubidus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-01-07 Classification: TRANSFERASE Ligands: A1EI6, SAH |
 |
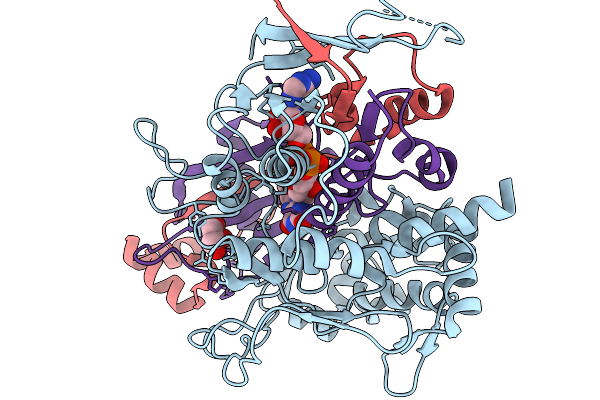
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
 |
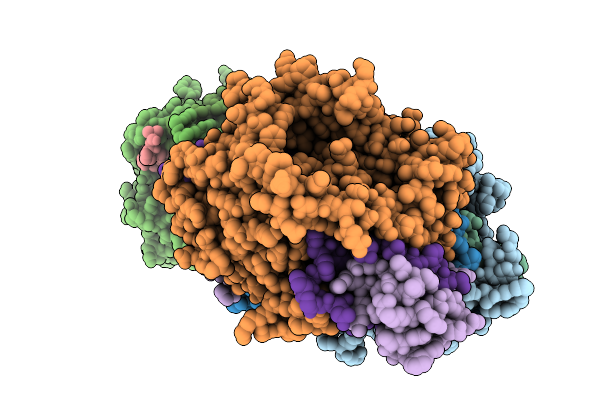
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GOL |
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD, GLU |
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:3.27 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Streptomyces sp. x-119-6
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: FAD |

