Search Count: 617
All
Selected
 |
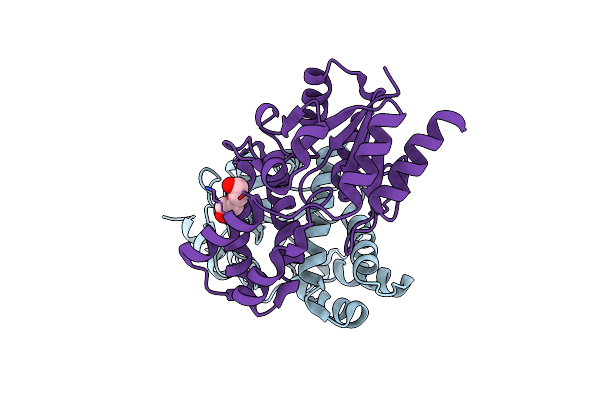
Ang-1 Domain Of The Hupe/Urej-2 Protein From Rhodobacteraceae Bacterium Rbang-1A With Cu Bound
Organism: Rhodobacteraceae bacterium pd-2
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-09-10 Classification: METAL BINDING PROTEIN Ligands: ZN, CU |
 |
Organism: Rhodobacteraceae bacterium qy30, Dna molecule
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-05-07 Classification: ANTIVIRAL PROTEIN/DNA Ligands: MN |
 |
Organism: Rhodobacteraceae bacterium qy30
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2025-05-07 Classification: ANTIVIRAL PROTEIN/DNA Ligands: PO4 |
 |
Organism: Rhodobacteraceae bacterium qy30
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2025-05-07 Classification: ANTIVIRAL PROTEIN Ligands: A1A3G, F2A, MN, A1A3H, D5M |
 |
Organism: Rhodobacteraceae bacterium qy30, Dna molecule
Method: ELECTRON MICROSCOPY Release Date: 2025-05-07 Classification: ANTIVIRAL PROTEIN |
 |
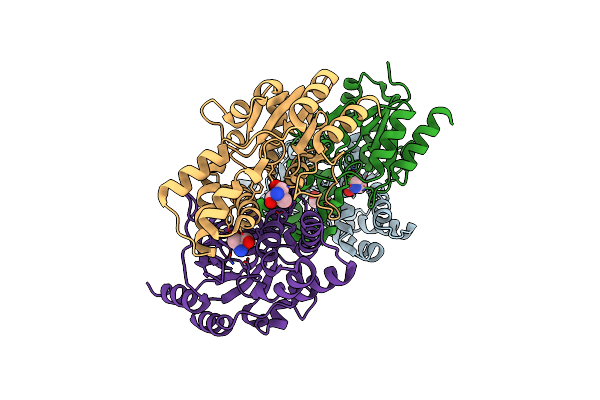
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2015-02-18 Classification: HYDROLASE Ligands: PEG |
 |
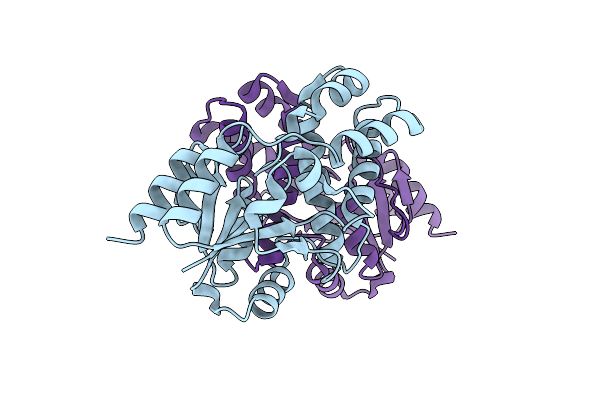
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2014-11-26 Classification: HYDROLASE Ligands: TRS |
 |
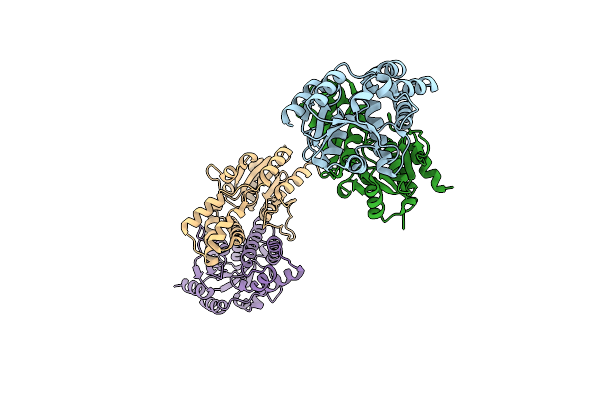
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2014-11-26 Classification: HYDROLASE |
 |
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2014-11-26 Classification: HYDROLASE |
 |
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2014-11-19 Classification: HYDROLASE |
 |
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2014-05-14 Classification: HYDROLASE Ligands: CL |
 |
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2014-05-14 Classification: HYDROLASE Ligands: CL, PGE, HE2 |
 |
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2013-05-01 Classification: HYDROLASE |
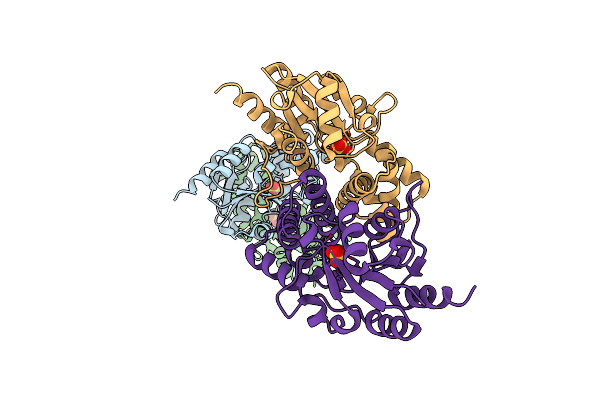 |
Sulfate Bound L-Haloacid Dehalogenase From A Rhodobacteraceae Family Bacterium
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2013-05-01 Classification: HYDROLASE Ligands: SO4 |
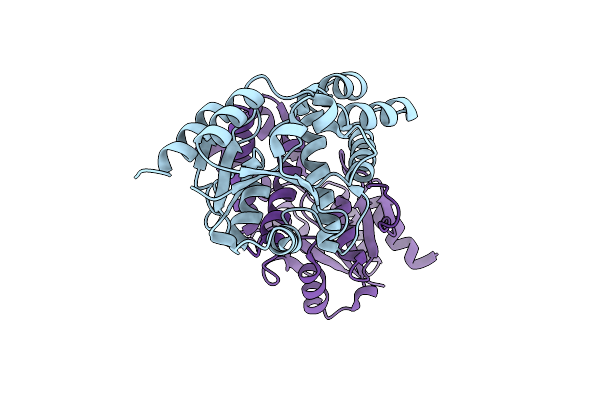 |
Chloroacetic Acid Complex Bound L-Haloacid Dehalogenase From A Rhodobacteraceae Family Bacterium
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2013-05-01 Classification: HYDROLASE |
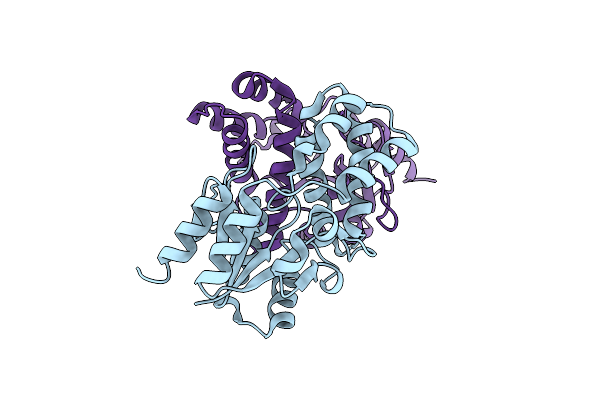 |
Chloropropionic Acid Complex Bound L-Haloacid Dehalogenase From A Rhodobacteraceae Family Bacterium
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2013-05-01 Classification: HYDROLASE |
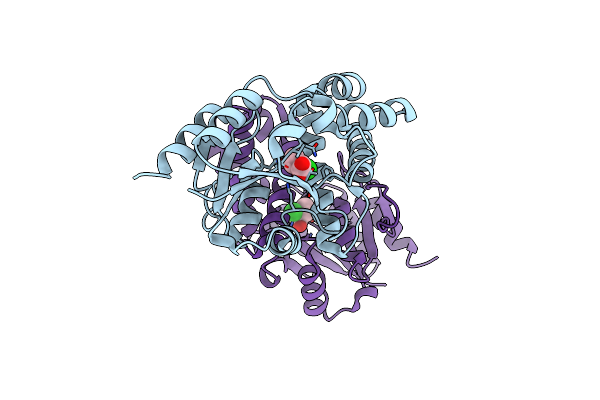 |
L-2-Chlorobutryic Acid Bound Complex L-Haloacid Dehalogenase From A Rhodobacteraceae Family Bacterium
Organism: Rhodobacteraceae
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2013-05-01 Classification: HYDROLASE Ligands: 39J |
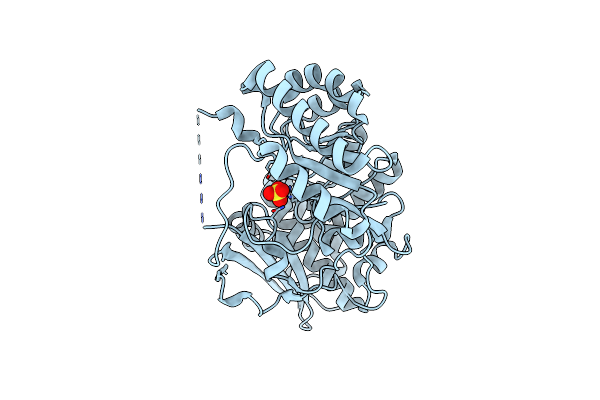 |
The Crystal Structure Of Mandelate Racemase/Muconate Lactonizing Enzyme From Rhodobacteraceae Bacterium
Organism: Rhodobacteraceae bacterium klh11
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2011-08-17 Classification: METAL BINDING PROTEIN Ligands: SO4 |
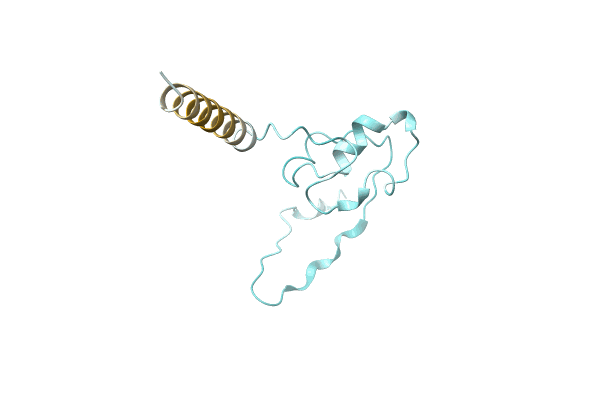 |
Organism: Rhodobacteraceae bacterium GWE1_64_9
Method: Alphafold Release Date: Classification: NA Ligands: NA |
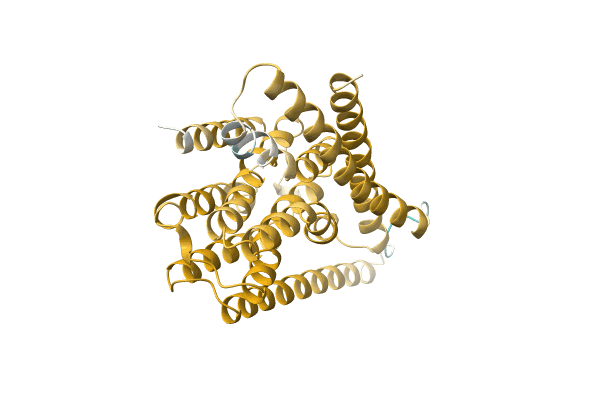 |
Organism: Rhodobacteraceae bacterium GWE1_64_9
Method: Alphafold Release Date: Classification: NA Ligands: NA |

