Search Count: 2,540
All
Selected
 |
The 100-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 1, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 150-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 2, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 200-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 3, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 150-K Crystal Structure Of Cyp199A4 Bound To 4-Methoxybenzoic Acid (Dataset 2, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: ANN, HEM, CL |
 |
The 200-K Crystal Structure Of Cyp199A4 Bound To 4-Methoxybenzoic Acid (Dataset 3, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.39 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: ANN, HEM, CL |
 |
The 300-K Crystal Structure Of Cyp199A4 Bound To 4-Methoxybenzoic Acid (Dataset 4, Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: HEM, ANN |
 |
The 200-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 1; Decreasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 150-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 2; Decreasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 100-K Crystal Structure Of Cyp199A4 Bound To 3-Methylaminobenzoic Acid (Dataset 3; Decreasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: OW4, HEM, CL |
 |
The 100-K Crystal Structure Of Cyp199A4 Bound To 4-Phenoxybenzoic Acid (Dataset 1; Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: HEM, JO4, CL |
 |
The 150-K Crystal Structure Of Cyp199A4 Bound To 4-Phenoxybenzoic Acid (Dataset 2; Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: JO4, HEM, CL |
 |
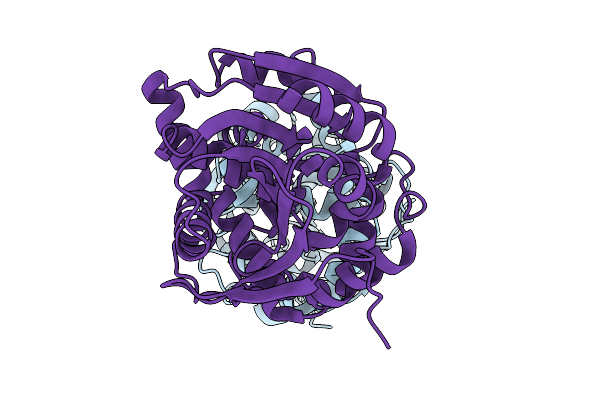
The 200-K Crystal Structure Of Cyp199A4 Bound To 4-Phenoxybenzoic Acid (Dataset 3; Increasing Temperature Series)
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: JO4, HEM, CL |
 |
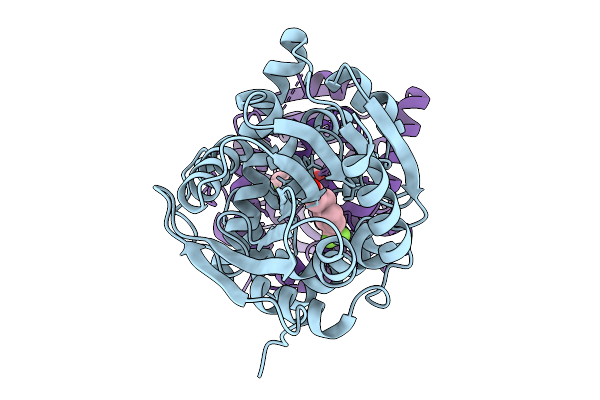
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Met/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-01-21 Classification: TRANSFERASE |
 |
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163-His280Ala With (S)-2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-01-07 Classification: HYDROLASE |
 |
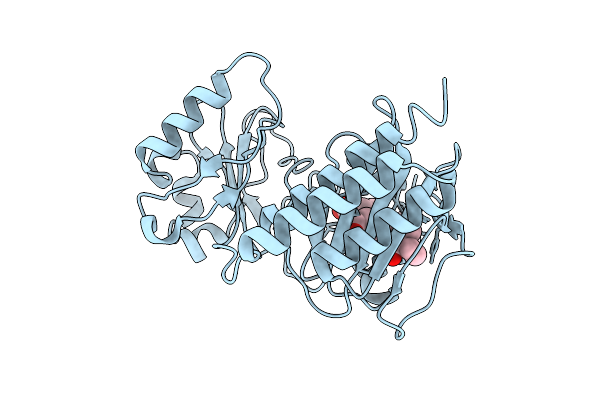
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Lys/Trp185Tyr/His280Ala With (S)-2-Fluoro-3-(4-(Trifluoromethyl)Phenyl)Propanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: A1ES3 |
 |
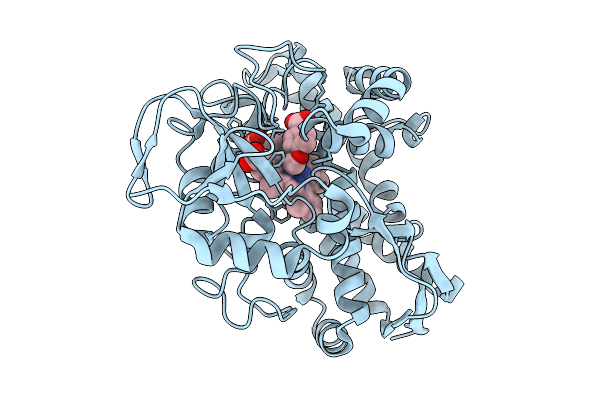
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Ser/Trp185Gly/His280Ala With (S)-2-Fluoro-3-(4-(Trifluoromethyl)Phenyl)Propanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: A1ES3 |
 |
Organism: Rhodopseudomonas palustris
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2025-10-29 Classification: FLUORESCENT PROTEIN Ligands: LBV |
 |
The High-Resolution Crystal Structure Of Wt Cyp199A4 Bound To 4-Methoxybenzoic Acid At 100 K
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2025-09-24 Classification: OXIDOREDUCTASE Ligands: HEM, ANN, CL |
 |
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: HEM, ANN, CL |
 |
Organism: Rhodopseudomonas palustris haa2
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: MW7, HEM, CL, ACT |

