Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Release Date: 2025-12-24 Classification: PLANT PROTEIN |
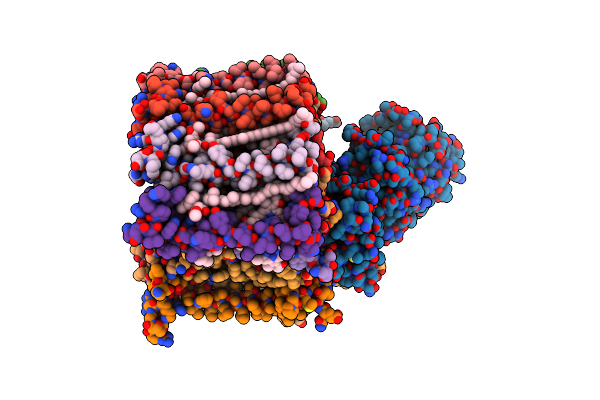 |
Organism: Chloroflexus aurantiacus j-10-fl
Method: ELECTRON MICROSCOPY Resolution:3.05 Å Release Date: 2025-02-19 Classification: PHOTOSYNTHESIS Ligands: BCL, U4Z, HEM, PGV, MQE, BPH, MN |
 |
Crystal Structure Of The Acvr1 (Alk2) Kinase Domain In Complex With Inhibitor Cdd-2789
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2024-11-27 Classification: TRANSFERASE Ligands: GOL, A1A29, PO4 |
 |
Crystal Structure Of The Acvr1 (Alk2) Kinase Domain In Complex With Inhibitor Cdd-2281
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2024-11-27 Classification: TRANSFERASE Ligands: PO4, GOL, A1A3C |
 |
Crystal Structure Of The Acvr1 (Alk2) Kinase Domain In Complex With Inhibitor Cdd-2282
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2024-11-27 Classification: TRANSFERASE Ligands: A1A3D, PO4 |
 |
Organism: Fittonia albivenis
Method: ELECTRON MICROSCOPY Release Date: 2024-07-10 Classification: PHOTOSYNTHESIS Ligands: CHL, CLA, LUT, XAT, BCR, LHG, LMG, HTG, PQN, SF4, LMT, DGD |
 |
Structure Of A Synthetic Circadian Clock Protein Kaic Mutant Of Cyanobacteria Synechococcus Elongatus Pcc 7942
Organism: Synechococcus elongatus (strain atcc 33912 / pcc 7942 / fachb-805)
Method: ELECTRON MICROSCOPY Release Date: 2024-06-12 Classification: BIOSYNTHETIC PROTEIN Ligands: ATP, MG, ADP |
 |
Crystal Structure Of Serine/Threonine-Protein Kinase 33 (Stk33) Kinase Domain In Complex With Inhibitor Cdd-2211
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-05 Classification: CYTOSOLIC PROTEIN Ligands: A1AAY |
 |
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-01-03 Classification: HYDROLASE/INHIBITOR Ligands: WIP, BCT |
 |
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2024-01-03 Classification: HYDROLASE/INHIBITOR Ligands: BCT, X6Q |
 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A-T268V In Complex With Pyd-N-C4-Phe And Hydroxylamine
Organism: Priestia megaterium nbrc 15308 = atcc 14581
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2023-12-27 Classification: OXIDOREDUCTASE Ligands: HEM, HOA, LXO, PHE |
 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A-T268V In Complex With Pyd-Pid-Phe And Hydroxylamine
Organism: Priestia megaterium nbrc 15308 = atcc 14581
Method: X-RAY DIFFRACTION Resolution:2.64 Å Release Date: 2023-12-27 Classification: OXIDOREDUCTASE Ligands: HOA, HEM, UCH, PHE |
 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A In Complex With Im-C6-Tyr-Tyr
Organism: Priestia megaterium, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: HEM |
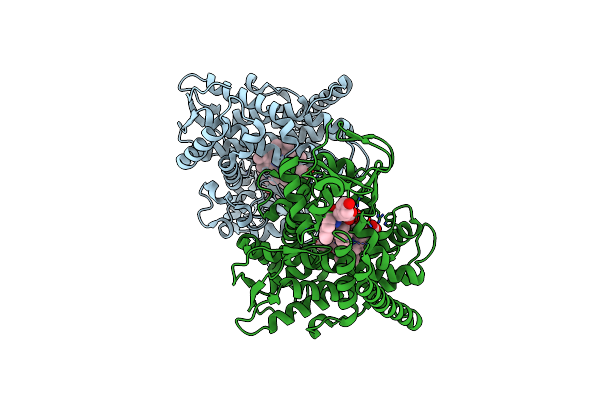 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A In Complex With Im-C6-Nap-Tyr
Organism: Priestia megaterium, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: HEM |
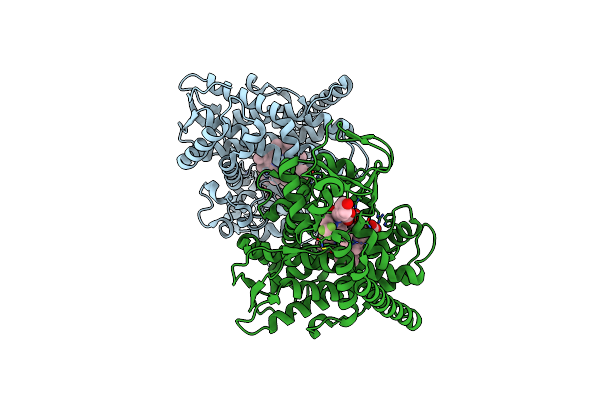 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A In Complex With Im-C6-Phe(4Cf3)-Tyr
Organism: Priestia megaterium, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: HEM |
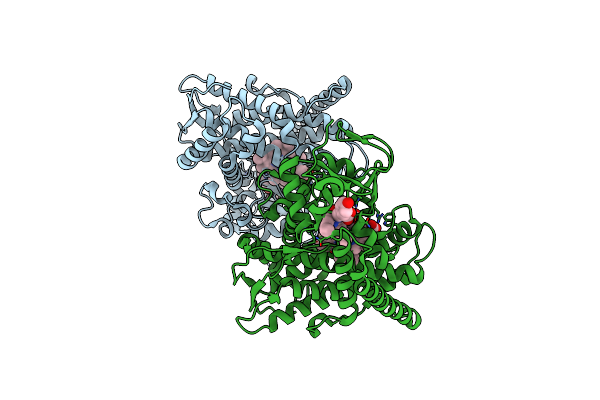 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A In Complex With Im-C6-Phe(4Ch3)-Tyr
Organism: Priestia megaterium, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: HEM |
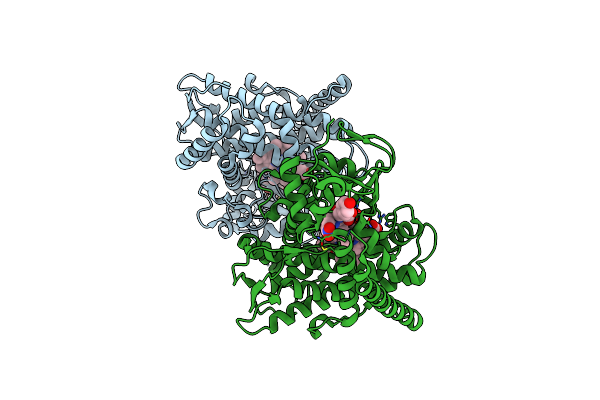 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A In Complex With Im-C6-Phe(4No2)-Tyr
Organism: Priestia megaterium, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: HEM |
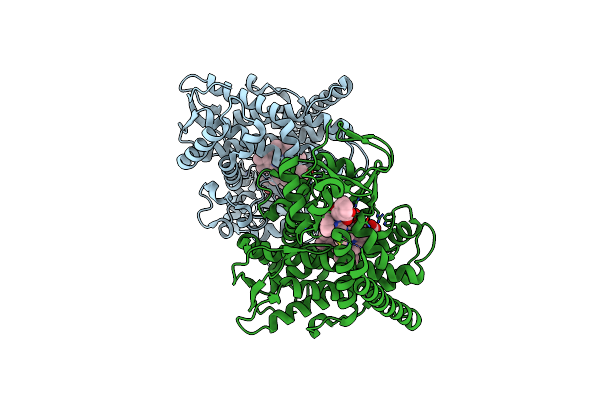 |
Crystal Structure Of The P450 Bm3 Heme Domain Mutant F87A-T268V In Complex With Im-N-C4-Phe-Phe
Organism: Priestia megaterium, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2023-12-13 Classification: OXIDOREDUCTASE Ligands: HEM |
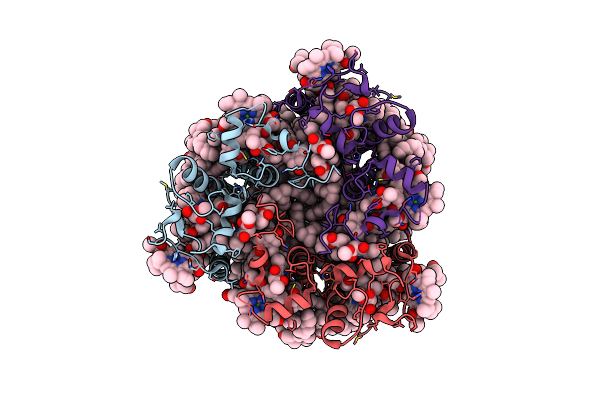 |
Organism: Bryopsis corticulans
Method: ELECTRON MICROSCOPY Release Date: 2023-09-06 Classification: PHOTOSYNTHESIS Ligands: 0UR, 0IE, NEX, CHL, CLA, LHG |
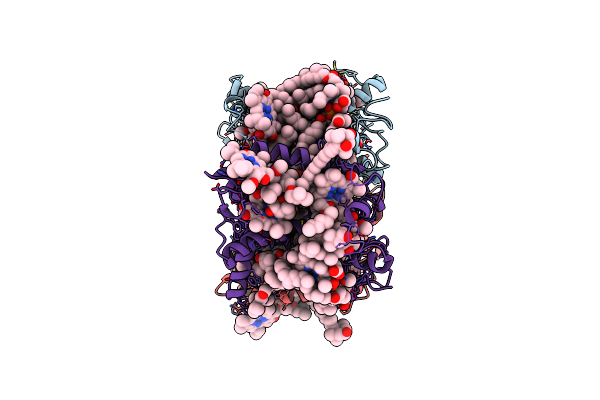 |
Organism: Bryopsis corticulans
Method: ELECTRON MICROSCOPY Release Date: 2023-09-06 Classification: PHOTOSYNTHESIS Ligands: 0UR, 0IE, NEX, CHL, CLA, LHG |