Search Count: 682
All
Selected
 |
Crystal Structure Of Chimeric Air Synthetase Constructed From E. Coli (Residues 1-168) And Pyrococcus Abyssi (Residues 163-334)
Organism: Escherichia coli, Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: LIGASE |
 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 43-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE Ligands: SO4 |
 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With An N-Terminal 54-Residues Deletion
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-02-11 Classification: LIGASE |
 |
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4 |
 |
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4, ADP |
 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase Bound To Adp And Mg(Ii)
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4, ADP, MG |
 |
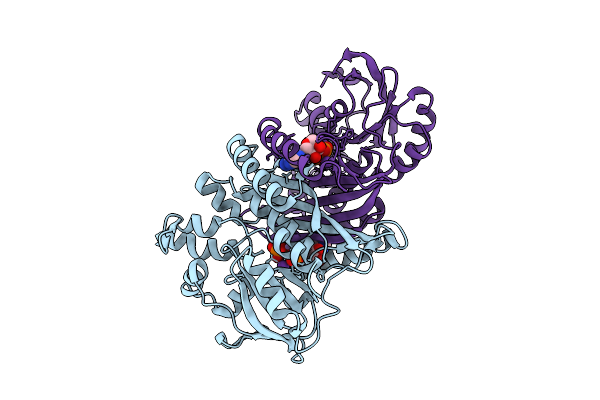
Crystal Structure Of Pyrococcus Abyssi Air Synthetase H239A Mutant Bound To Amppnp
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: ANP, MN, K |
 |
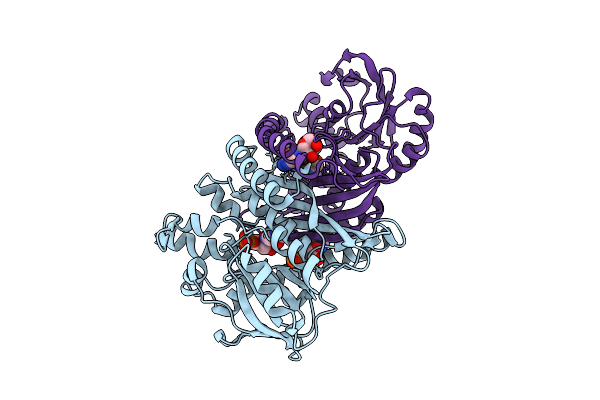
Crystal Structure Of Pyrococcus Abyssi Air Synthetase H239A Mutant Bound To Adp And Pi
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: ADP, PO4, MN, K |
 |
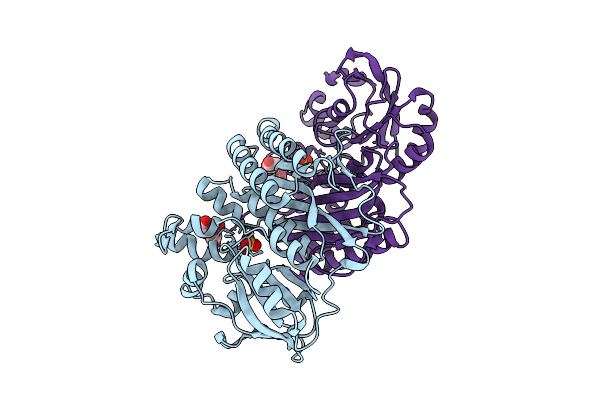
Crystal Structure Of Pyrococcus Abyssi Air Synthetase N186A Mutant Bound To Fgar And Amp
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: FGR, AMP |
 |
Crystal Structure Of Pyrococcus Abyssi Air Synthetase With Residues 51-54 Replaced By Neisseria Gonorrhoeae Residues 49-56
Organism: Pyrococcus abyssi ge5
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-10-15 Classification: LIGASE Ligands: SO4, FLC |
 |
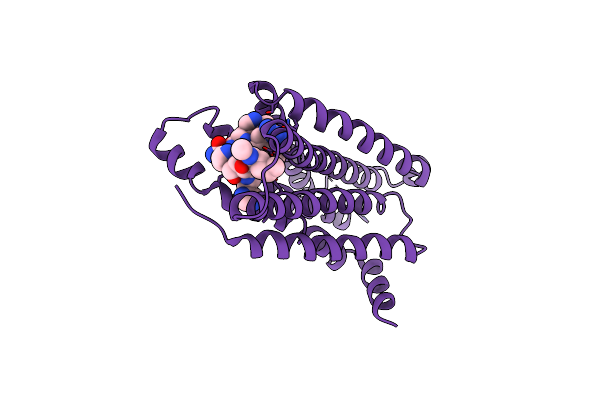
Organism: Homo sapiens, Pyrococcus abyssi
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: 5YM, A1EM1, 1DO, A1EM3, A1EQ8, A1EM2 |
 |
Organism: Escherichia coli, Homo sapiens, Pyrococcus abyssi ge5, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2025-02-26 Classification: MEMBRANE PROTEIN Ligands: CA |
 |
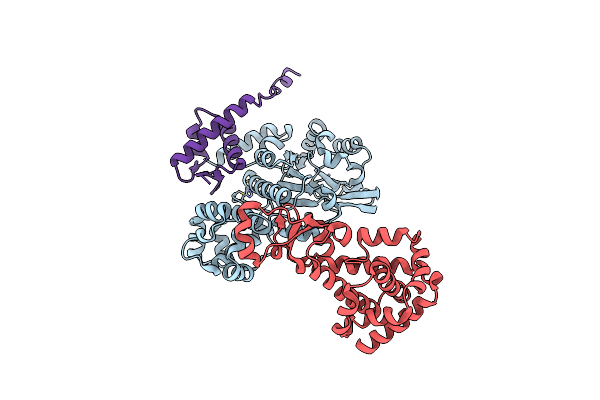
Cryo-Em Structure Of Saccharolobus Solfataricus 30S Initiation Complex Bound To Sd Mrna With H44 In Up Position
Organism: Escherichia coli, Pyrococcus abyssi, Saccharolobus solfataricus p2
Method: ELECTRON MICROSCOPY Release Date: 2025-01-15 Classification: TRANSLATION Ligands: MG, SPM, ZN |
 |
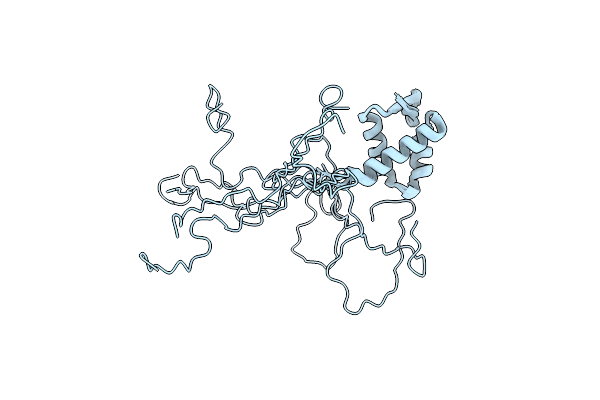
Organism: Pyrococcus abyssi
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2024-12-04 Classification: DNA BINDING PROTEIN Ligands: ZN |
 |
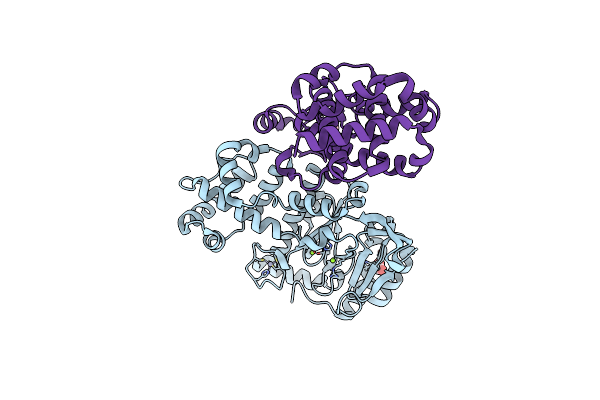
Organism: Pyrococcus abyssi ge5
Method: SOLUTION NMR Release Date: 2024-12-04 Classification: REPLICATION |
 |
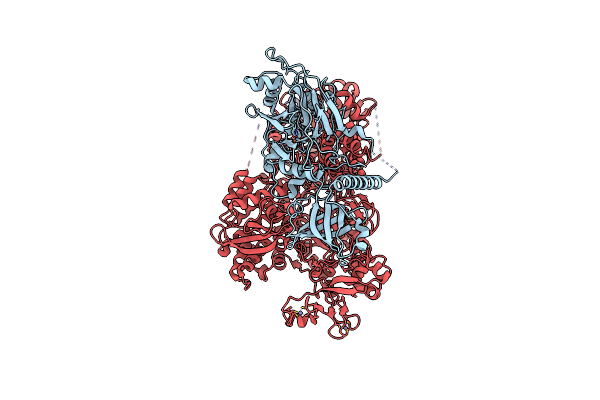
Crystal Structure Of The Heterodimeric Primase From Pyrococcus Abyssi (Deletion Of The Pril-Ctd Domain)
Organism: Pyrococcus abyssi
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2024-12-04 Classification: REPLICATION Ligands: ZN, MG, FMT |
 |
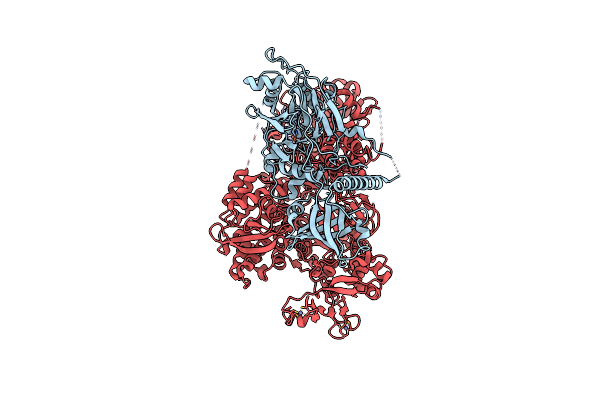
Pyrococcus Abyssi Pold In Complex With Rpa2 Winged-Helix Domain Class 1 (Composite Map)
Organism: Pyrococcus abyssi ge5
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: REPLICATION Ligands: FE, ZN |
 |
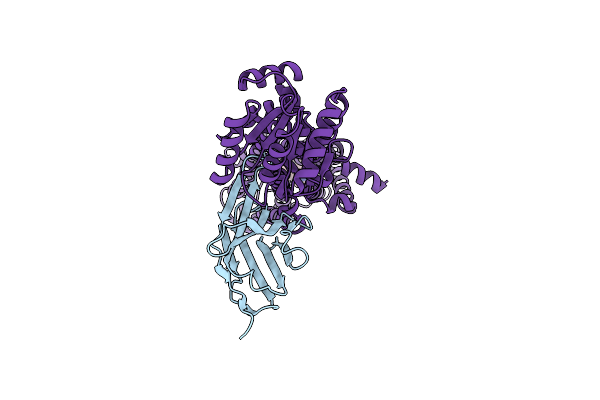
Pyrococcus Abyssi Pold In Complex With Rpa2 Winged-Helix Domain Class 2 (Composite Map)
Organism: Pyrococcus abyssi ge5
Method: ELECTRON MICROSCOPY Release Date: 2024-12-04 Classification: REPLICATION Ligands: FE, ZN |
 |
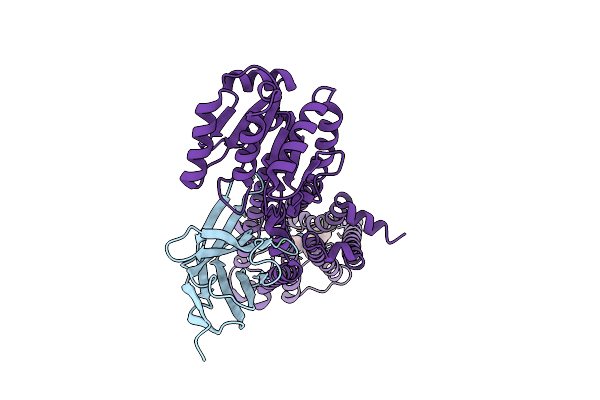
Structural Mechanism Of Cb1R Binding To Peripheral And Biased Inverse Agonists
Organism: Homo sapiens, Pyrococcus abyssi ge5, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2024-11-20 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: 7DY |
 |
Structural Mechanism Of Cb1R Binding To Peripheral And Biased Inverse Agonists
Organism: Lama glama, Homo sapiens, Pyrococcus abyssi ge5
Method: ELECTRON MICROSCOPY Release Date: 2024-11-20 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: A1AKO |

