Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
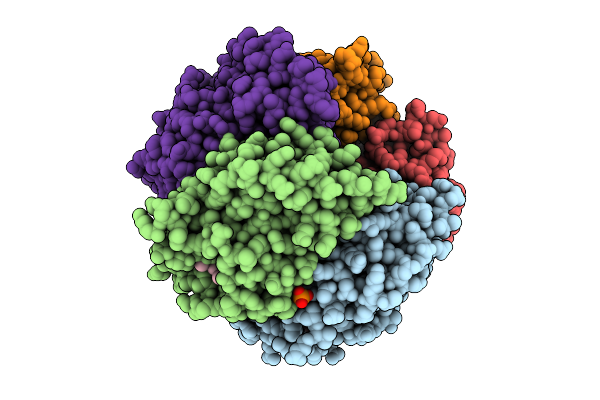 |
G169V Variant Of Streptomyces Coelicolor Coproheme Decarboxylase In Complex With Monovinyl, Monopropionate Deuteroheme
Organism: Streptomyces coelicolor
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: BIOSYNTHETIC PROTEIN Ligands: VOV, PO4, NA |
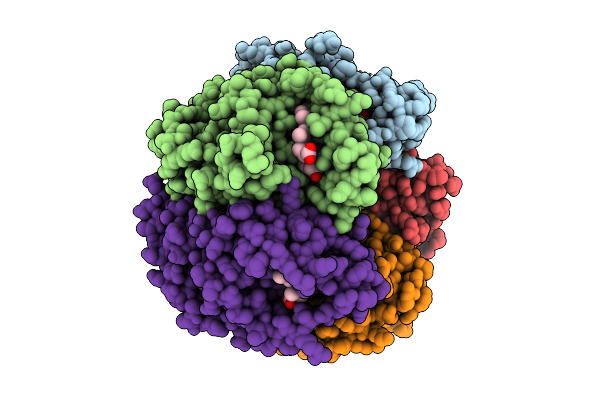 |
G169L Variant Of Coproheme Decarboxylase From Streptomyces Coelicolor In Complex With Monovinyl, Monopropionate Deuteroheme
Organism: Streptomyces coelicolor
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: BIOSYNTHETIC PROTEIN Ligands: VOV |
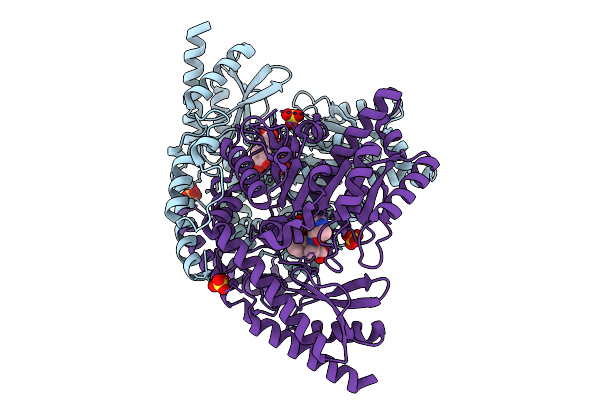 |
Re-Refined Of Crystal Structure Of Dopa Decarboxylase In Complex With The Inhibitor Carbidopa (1Js3) With Ketoenamine Form Of Carbidopa
Organism: Sus scrofa
Method: X-RAY DIFFRACTION Release Date: 2025-09-10 Classification: LYASE Ligands: A1BCS, SO4 |
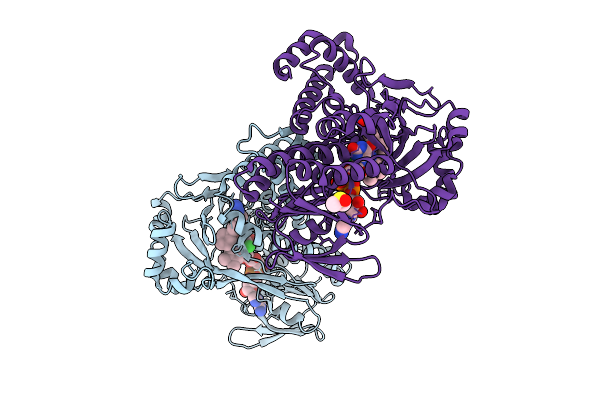 |
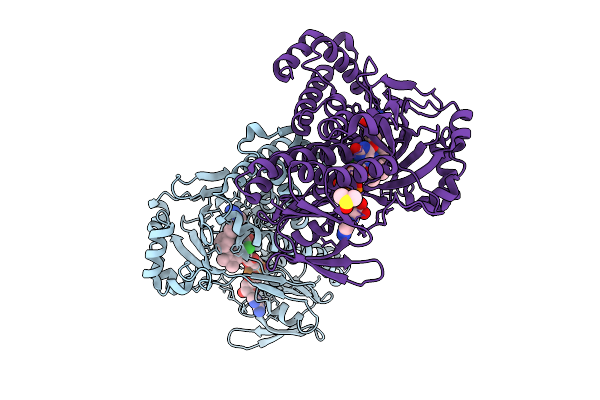
Kynurenine Monooxygenase From Pseudomonas Fluorescens Complexed With 4-Cyclohexylphenylacetyltetrazole
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Release Date: 2025-08-27 Classification: FLAVOPROTEIN/INHIBITOR Ligands: A1AIN, FAD, CL, DMS |
 |
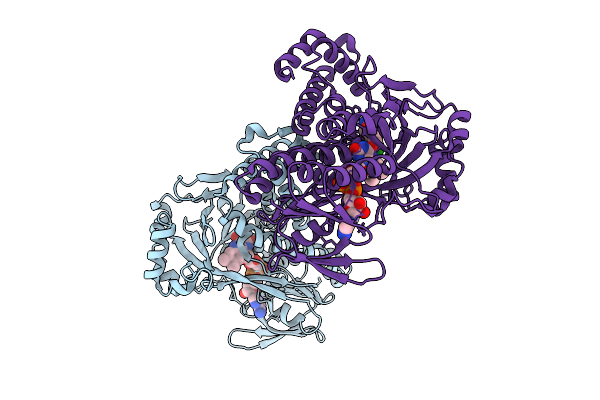
Kynurenine Monooxygenase From Pseudomonas Fluorescens Complexed With Biphenylacetyltetrazole
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Release Date: 2025-08-27 Classification: FLAVOPROTEIN/INHIBITOR Ligands: FAD, CL, A1AIU, DMS |
 |
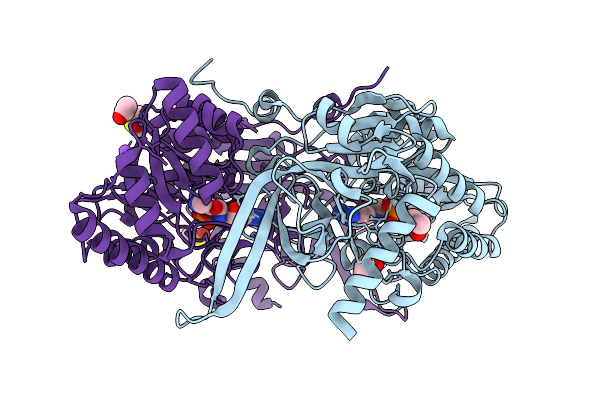
Kynurenine Monooxygenase From Pseudomonas Fluorescens Complexed With 4-Chlorophenylacetyltetrazole
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Release Date: 2025-08-27 Classification: FLAVOPROTEIN/INHIBITOR Ligands: FAD, CL, A1AI5 |
 |
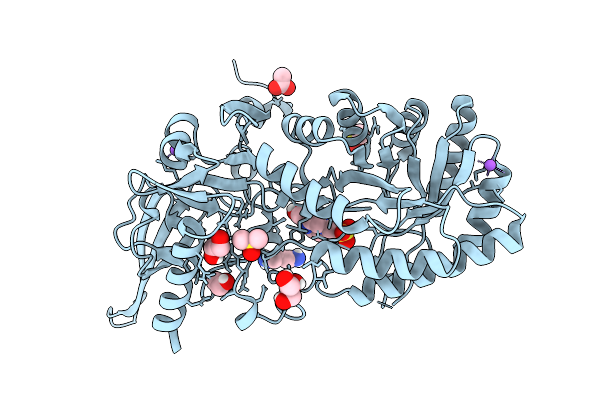
C387S Variant Of D-Ornithine/D-Lysine Decarboxylase Complexed With Pmp And 4-Guanidinobutanal
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Release Date: 2025-08-06 Classification: LYASE Ligands: DMS, NA, PMP, A1AZK, ACT, CL |
 |
C387S Variant Of D-Ornithine/D-Lysine Decarboxylase Complexed With Hepes And Putrescine
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Release Date: 2025-08-06 Classification: LYASE Ligands: CL, NA, PUT, EPE, DMS, ACT, EDO, GOL |
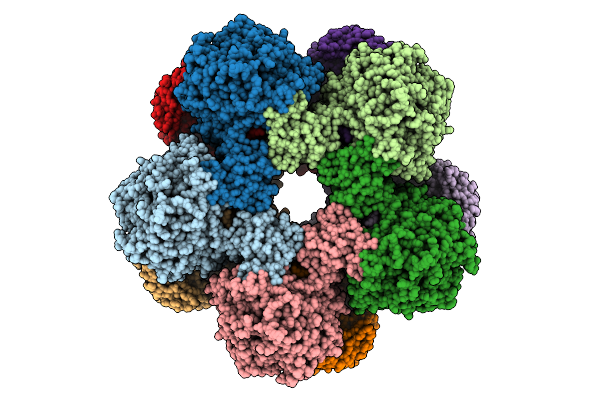 |
Cryoem Structure Of Inducible Lysine Decarboxylase From Hafnia Alvei D-Hydrazino-Lysine Analog At 2.3 Angstrom Resolution
Organism: Hafnia alvei atcc 51873
Method: ELECTRON MICROSCOPY Release Date: 2025-07-30 Classification: LYASE Ligands: A1BD0 |
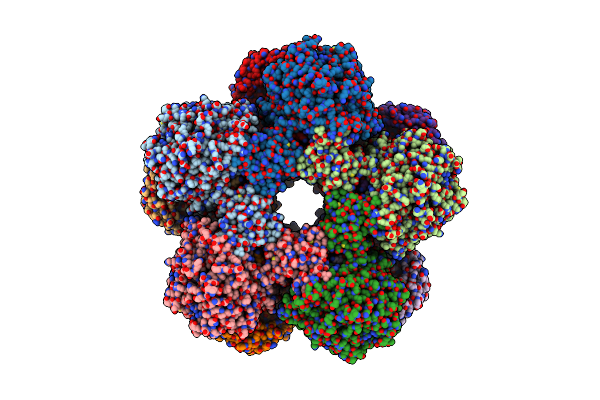 |
Cryoem Structure Of Holoenzyme Of Inducible Lysine Decarboxylase From Hafnia Alvei Holoenzyme At 2.19 Angstrom Resolution
Organism: Hafnia alvei atcc 51873
Method: ELECTRON MICROSCOPY Release Date: 2025-06-04 Classification: LYASE |
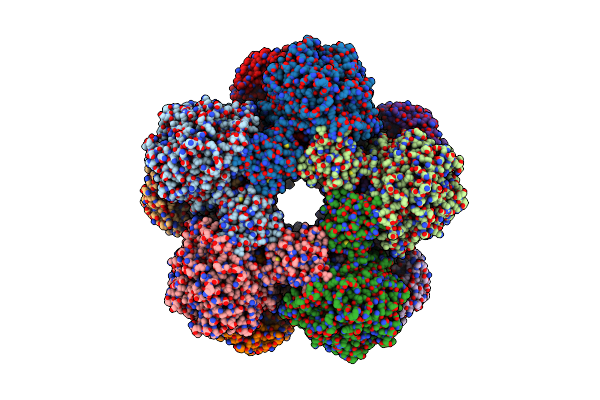 |
Cryoem Structure Of Inducible Lysine Decarboxylase From Hafnia Alvei L-Hydrazino-Lysine Analog At 2.04 Angstrom Resolution
Organism: Hafnia alvei atcc 51873
Method: ELECTRON MICROSCOPY Release Date: 2025-06-04 Classification: LYASE Ligands: A1BD1 |
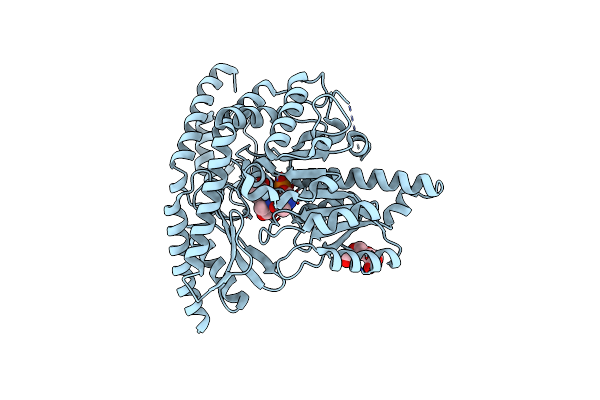 |
X-Ray Structure Of Human Holo Aromatic L-Amino Acid Decarboxylase (Aadc) Complex With Carbidopa At Physiological Ph
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: BIOSYNTHETIC PROTEIN Ligands: PLP, 142, PG4 |
 |
Proteus Vulgaris Tryptophan Indole-Lyase Complexed With L-Ethionine And Na+
Organism: Proteus vulgaris
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2025-04-16 Classification: LYASE Ligands: DMS, EDO, ESC, NA, YAR, YAE |
 |
Proteus Vulgaris Tryptophan Indole-Lyase Complexed With L-Trp And Benzimidazole
Organism: Proteus vulgaris
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2025-02-12 Classification: LYASE Ligands: K, BZI, 0JO, DMS, PLT, A1A2Z |
 |
Room-Temperature X-Ray Structure Of Thermus Thermophilus Serine Hydroxymethyltransferase (Shmt) With Plp-Glycine External Aldimine And 5-Formyltetrahydrofolate (Folinic Acid)
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: SO4, A1AQW, FFO |
 |
X-Ray Structure Of Thermus Thermophilus Serine Hydroxymethyltransferase With Plp-L-Ser External Aldimine And 5-Formyltetrahydrofolate (Folinic Acid)
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: SER, SO4, KOU, FFO |
 |
Room-Temperature X-Ray Structure Of Human Mitochondrial Serine Hydroxymethyltransferase (Hshmt2) With Plp-Glycine External Aldimine And 5-Formyltetrahydrofolate (Folinic Acid)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: A1AQW, FFO |
 |
Joint X-Ray/Neutron Structure Of Thermus Thermophilus Serine Hydroxymethyltransferase (Tthshmt) In Internal Aldimine State And Folinic Acid Bound
Organism: Thermus thermophilus hb8
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.30 Å, 2.00 Å Release Date: 2024-08-28 Classification: TRANSFERASE Ligands: SO4, FFO, ACT, DOD |
 |
Proteus Vulgaris Tryptophan Indole-Lyase Aminoacrylate Complex With Benzimidazole
Organism: Proteus vulgaris
Method: X-RAY DIFFRACTION Resolution:1.51 Å Release Date: 2024-05-29 Classification: LYASE Ligands: K, BZI, DMS, 0JO, CL |
 |
Organism: Proteus vulgaris
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2024-01-31 Classification: LYASE Ligands: K, A1ACN, DMS, A1ABZ |