Search Count: 6,358
All
Selected
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-03-04 Classification: CYTOSOLIC PROTEIN |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-04 Classification: CYTOSOLIC PROTEIN Ligands: 1PE, ACT |
 |
Crystal Structure Of Sigma28/Flgm Complex From Pseudomonas Aeruginosa At 1.95 Angstrom Resolution
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-02-25 Classification: TRANSCRIPTION |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-02-18 Classification: MEMBRANE PROTEIN Ligands: A1IJR, EDO, C8E, GOL, FE, K |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-02-11 Classification: ANTIBIOTIC Ligands: A1I8B |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-02-11 Classification: TOXIN |
 |
Crystal Structure Of Anti-Crispr Acrie7 Determined By Experimental Mad Phasing With Hg
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-02-04 Classification: DNA BINDING PROTEIN Ligands: HG |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-02-04 Classification: UNKNOWN FUNCTION Ligands: CL |
 |
Crystal Structure Of Adp-Ribosylated Endolytic Muramidase Tdem From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa (strain atcc 15692 / dsm 22644 / cip 104116 / jcm 14847 / lmg 12228 / 1c / prs 101 / pao1)
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-21 Classification: DE NOVO PROTEIN Ligands: BEN, ZN |
 |
Organism: Pseudomonas aeruginosa pao1
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: LIGASE |
 |
Organism: Pseudomonas aeruginosa pao1
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: LIGASE |
 |
Organism: Pseudomonas aeruginosa pao1
Method: ELECTRON MICROSCOPY Release Date: 2026-01-07 Classification: LIGASE |
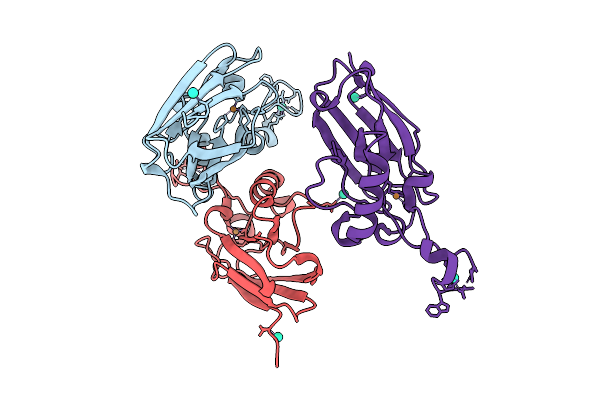 |
Organism: Pseudomonas aeruginosa (strain atcc 15692 / dsm 22644 / cip 104116 / jcm 14847 / lmg 12228 / 1c / prs 101 / pao1)
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2025-12-24 Classification: ELECTRON TRANSPORT Ligands: CU, TB |
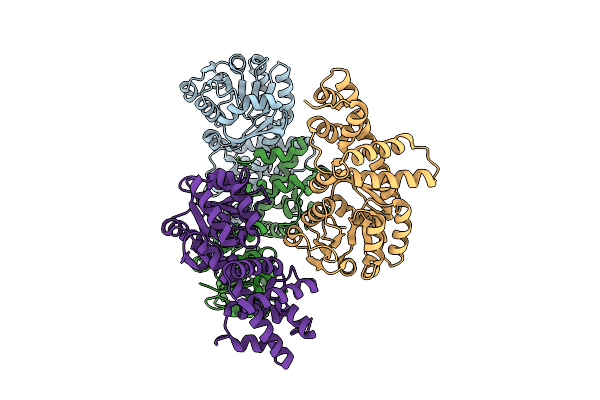 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2025-12-24 Classification: TRANSFERASE |
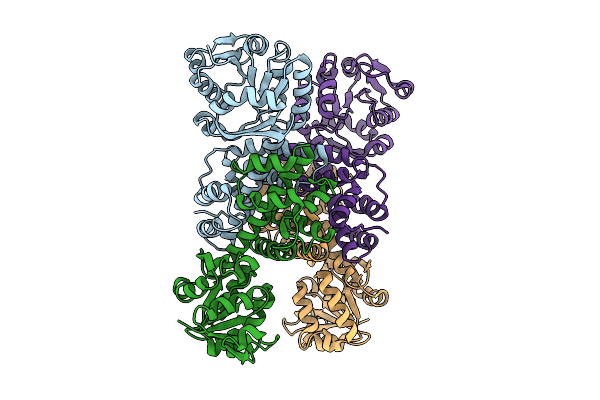 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2025-12-24 Classification: HYDROLASE |
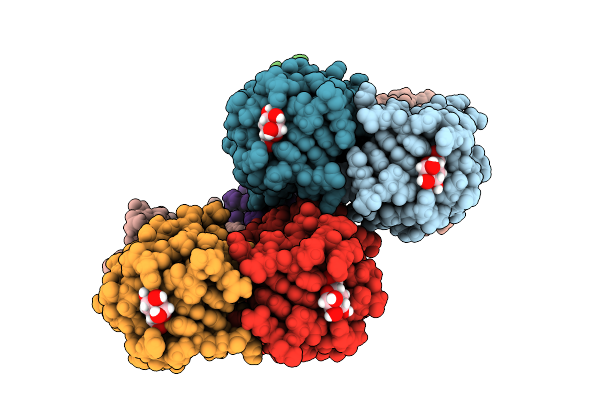 |
Joint X-Ray/Neutron Room Temperature Structure Of Perdeuterated Leca Lectin In Complex With Deuterated Galactose
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:1.49 Å, 1.9 Å Release Date: 2025-12-24 Classification: SUGAR BINDING PROTEIN Ligands: GLA, CA |
 |
Structure Of Flagellar Hook Subunit Flge D-I Domain In Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa (strain atcc 15692 / dsm 22644 / cip 104116 / jcm 14847 / lmg 12228 / 1c / prs 101 / pao1)
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: MOTOR PROTEIN |
 |
Structure Of Flagellar Hook Subunit Flge D-Ii Domain In Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: MOTOR PROTEIN |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2025-12-17 Classification: TRANSFERASE Ligands: RFP |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:4.54 Å Release Date: 2025-12-10 Classification: MOTOR PROTEIN |

