Search Count: 275
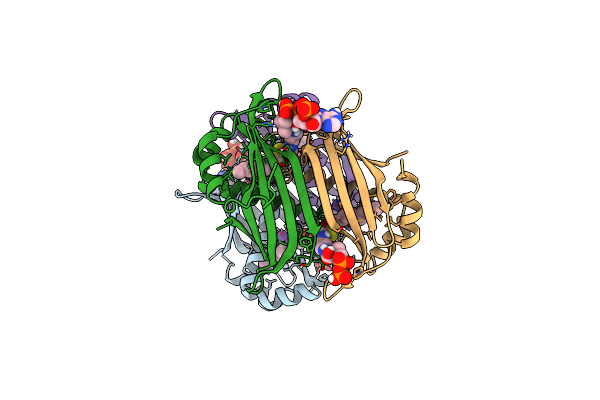 |
Crystal Structure Of The Espe7 Thioesterase Mutant R35A From The Esperamicin Biosynthetic Pathway At 1.6 A
Organism: Actinomadura verrucosospora
Method: X-RAY DIFFRACTION Release Date: 2025-06-25 Classification: HYDROLASE Ligands: K, DCC |
 |
Organism: Bdellovibrio bacteriovorus hd100
Method: X-RAY DIFFRACTION Resolution:2.17 Å Release Date: 2023-10-25 Classification: UNKNOWN FUNCTION Ligands: GOL, EDO |
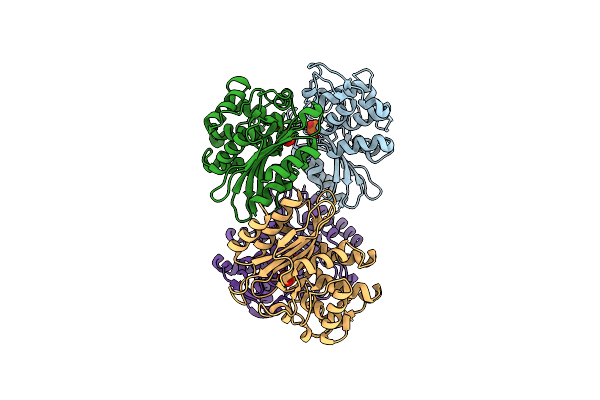 |
Xfel Structure Of Mycobacterium Tuberculosis Beta Lactamase Microcrystals Mixed With Sulbactam For 3 Ms
Organism: Mycobacterium tuberculosis str. beijing/w bt1
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2023-09-20 Classification: HYDROLASE/Inhibitor Ligands: PO4 |
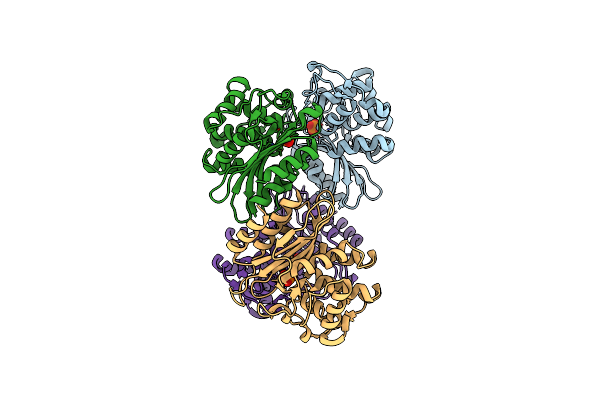 |
Xfel Structure Of Mycobacterium Tuberculosis Beta Lactamase Microcrystals Mixed With Sulbactam For 6 Ms
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2023-09-20 Classification: HYDROLASE/Inhibitor Ligands: PO4 |
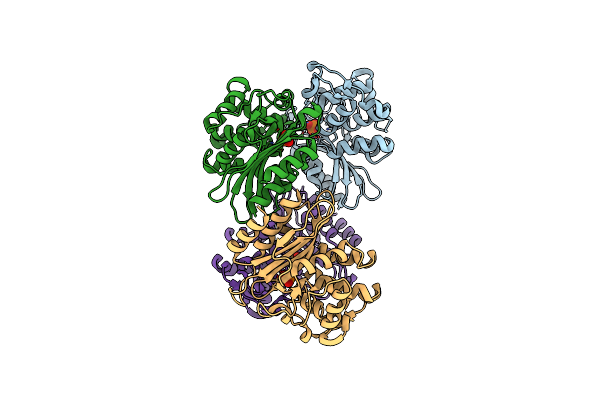 |
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2023-09-20 Classification: HYDROLASE/Inhibitor Ligands: PO4 |
 |
Xfel Structure Of Mycobacterium Tuberculosis Beta Lactamase Microcrystals Mixed With Sulbactam For 700 Ms
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2023-09-20 Classification: HYDROLASE/Inhibitor Ligands: PO4, TSL |
 |
Crystal Structure Of The N-Terminal Domain Of The Cryptic Surface Protein (Cd630_25440) From Clostridium Difficile.
Organism: Clostridioides difficile 630
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-05-10 Classification: UNKNOWN FUNCTION Ligands: CL, EDO, GOL |
 |
Xfel Structure Of Beta Lactamase Microcrystals Mixed With Sulbactam Solution For 15Ms
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2023-03-08 Classification: HYDROLASE Ligands: PO4, TSL |
 |
Xfel Structure Of Mycobacterium Tuberculosis Beta Lactamase Microcrystals Mixed With Sulbactam For 30Ms
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2023-03-08 Classification: HYDROLASE/Inhibitor Ligands: PO4, 0RN, TSL |
 |
Xfel Structure Of Mycobacterium Tuberculosis Beta Lactamase Microcrystals Mixed With Sulbactam For 240Ms
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2023-03-08 Classification: HYDROLASE/Inhibitor Ligands: PO4, TSL |
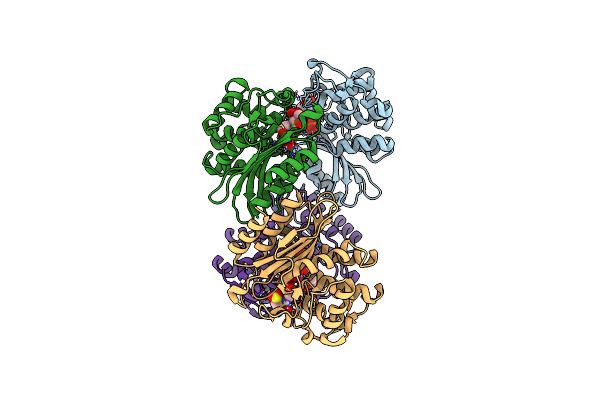 |
Cryo Structure Of Mycobacterium Tuberculosis Beta Lactamase Microcrystals Mixed With Sulbactam For 3 Hours
Organism: Mycobacterium tuberculosis
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2023-03-08 Classification: HYDROLASE/Inhibitor Ligands: PO4, TSL |
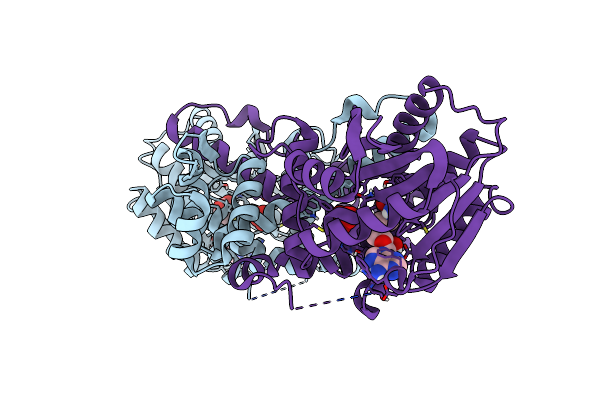 |
Crystal Structure Of Mfng, An L- And D-Tyrosine O-Methyltransferase From The Marformycin Biosynthesis Pathway Of Streptomyces Drozdowiczii, With Sah Bound At 1.35 A Resolution (P212121 - Form I)
Organism: Streptomyces drozdowiczii
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2022-10-12 Classification: TRANSFERASE Ligands: SAH, UNL |
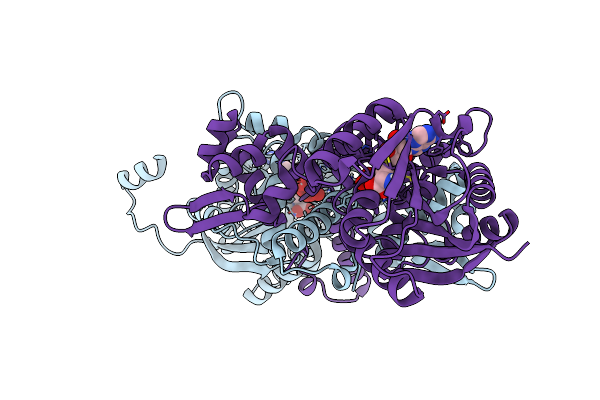 |
Crystal Structure Of Mfng, An L- And D-Tyrosine O-Methyltransferase From The Marformycin Biosynthesis Pathway Of Streptomyces Drozdowiczii, With Sah Bound At 1.2 A Resolution (P212121 - Form Ii)
Organism: Streptomyces drozdowiczii
Method: X-RAY DIFFRACTION Resolution:1.14 Å Release Date: 2022-10-12 Classification: TRANSFERASE Ligands: SAH, UNL |
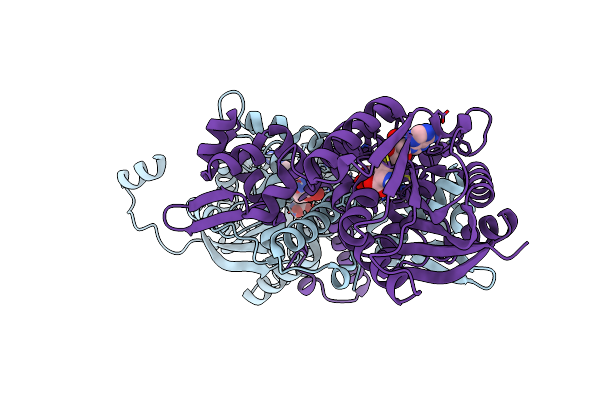 |
Crystal Structure Of Mfng, An L- And D-Tyrosine O-Methyltransferase From The Marformycin Biosynthesis Pathway Of Streptomyces Drozdowiczii, With Sah And L-Tyrosine Bound At 1.4 A Resolution (P212121 - Form Ii)
Organism: Streptomyces drozdowiczii
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2022-09-28 Classification: TRANSFERASE Ligands: SAH, TYR, UNL |
 |
Crystal Structure Of Ttnm, A Fe(Ii)-Alpha-Ketoglutarate-Dependent Hydroxylase From The Tautomycetin Biosynthesis Pathway In Streptomyces Griseochromogenes At 2 A.
Organism: Streptomyces griseochromogenes
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2022-07-06 Classification: OXIDOREDUCTASE Ligands: FE2, CL |
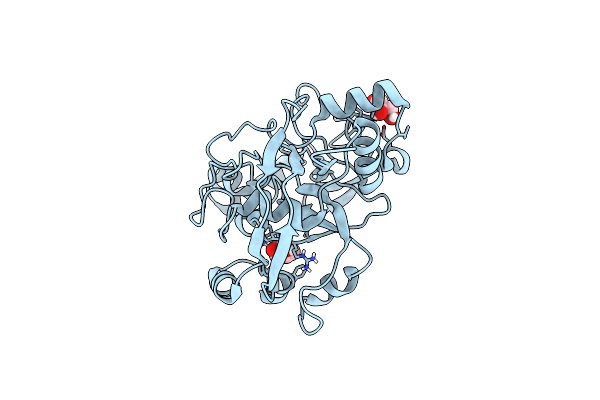 |
Structure Of Calu17 From The Calicheamicin Biosynthesis Pathway Of Micromonospora Echinospora
Organism: Micromonospora echinospora
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2021-07-28 Classification: UNKNOWN FUNCTION Ligands: GOL |
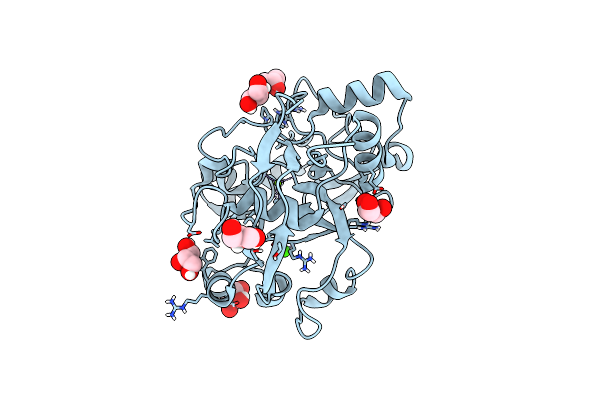 |
Structure Of Calu17 From The Calicheamicin Biosynthesis Pathway Of Micromonospora Echinospora
Organism: Micromonospora echinospora
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2021-07-28 Classification: UNKNOWN FUNCTION Ligands: CA, GOL, MG, CL |
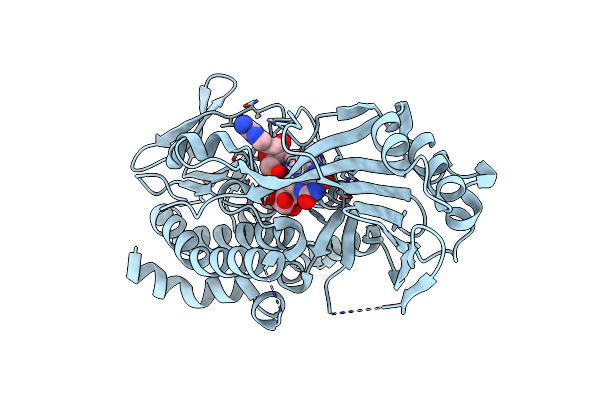 |
Organism: Penicillium citrinum
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2021-06-16 Classification: BIOSYNTHETIC PROTEIN Ligands: FAD |
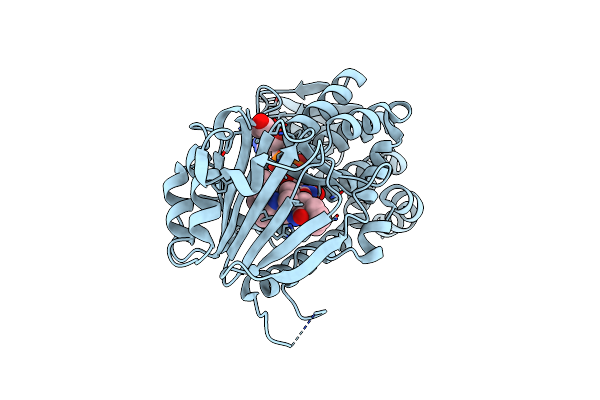 |
Organism: Penicillium citrinum
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2021-06-16 Classification: BIOSYNTHETIC PROTEIN Ligands: FAD, WU4, EDO, CL |
 |
The Crystal Structure Of A Collagen Galactosylhydroxylysyl Glucosyltransferase From Human
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2021-04-14 Classification: TRANSFERASE Ligands: MN, MG, UDP, TRS |

