Search Count: 166
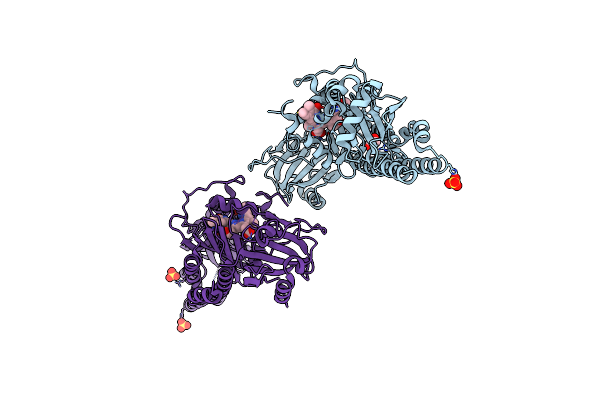 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State (I0A).
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL, EDO |
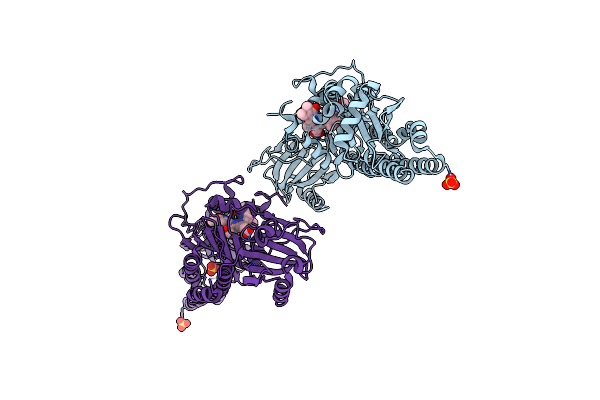 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In Its Pfr State (I0B).
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4 |
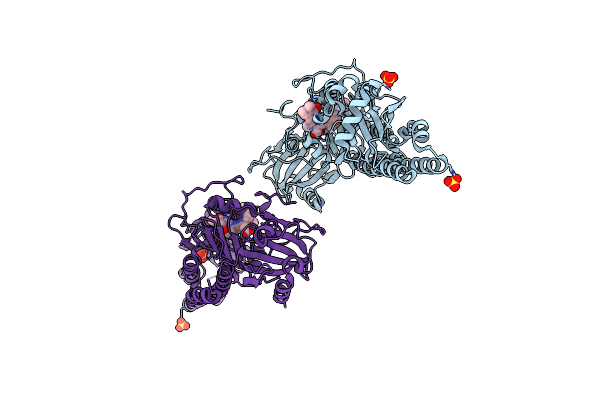 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I1 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4 |
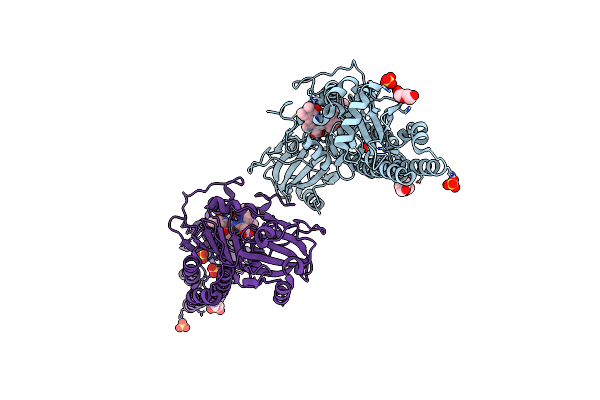 |
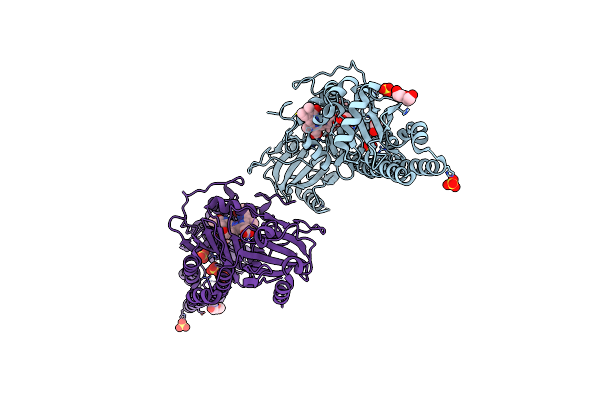
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I2 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, PGE, PEG, CL |
 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I3 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, GOL, PEG |
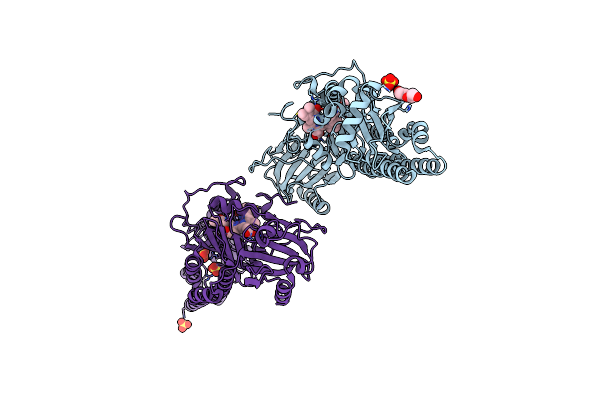 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I4 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL, PEG |
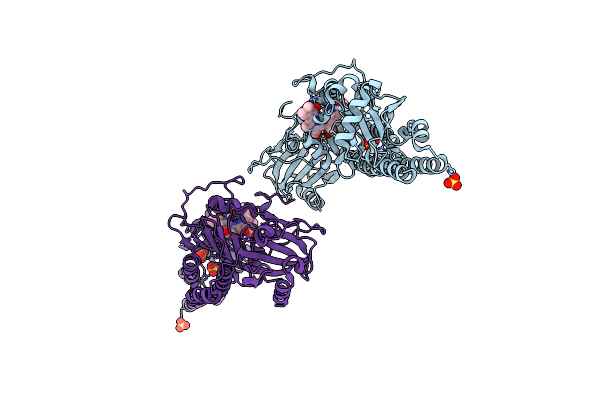 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I5 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL |
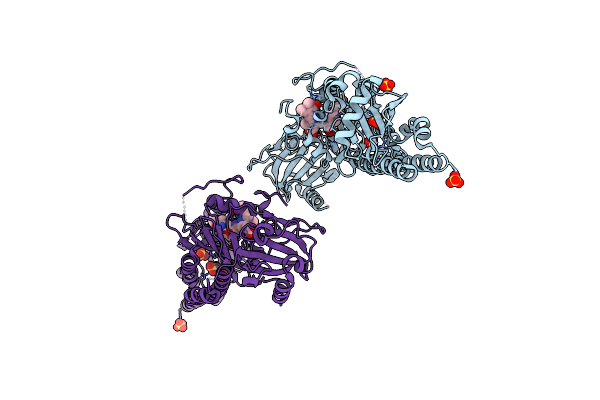 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I6 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, CL |
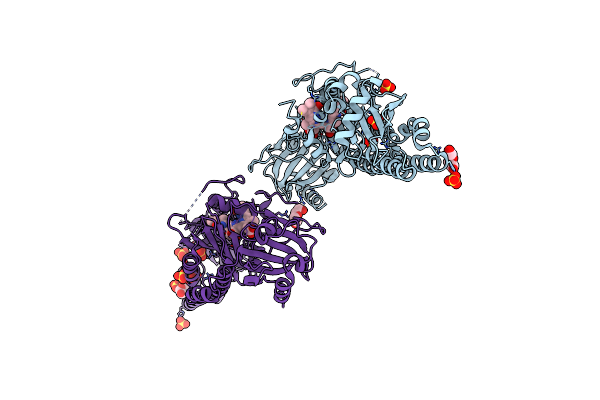 |
Serial Femtosecond X-Ray Structure Of A Fluorescence Optimized Bathy Phytochrome Pairfp2 Derived From Wild-Type Agp2 In I7 Intermediate State.
Organism: Agrobacterium fabrum str. c58
Method: X-RAY DIFFRACTION Release Date: 2025-05-14 Classification: SIGNALING PROTEIN Ligands: EL5, SO4, PEG |
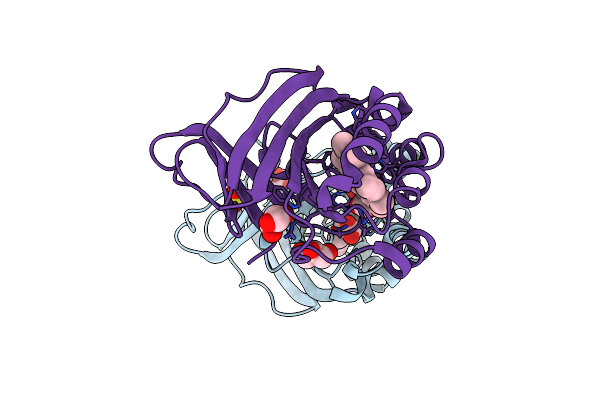 |
Crystal Structure Of Chlorite Dismutase At 3000 Ev Based On Analytical Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-07-03 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |
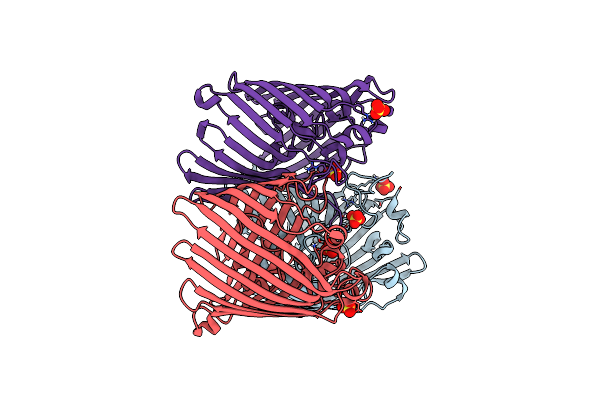 |
Crystal Structure Of Ompk36 Gd At 3500 Ev Based On Spherical Harmonics Absorption Corrections
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN Ligands: SO4 |
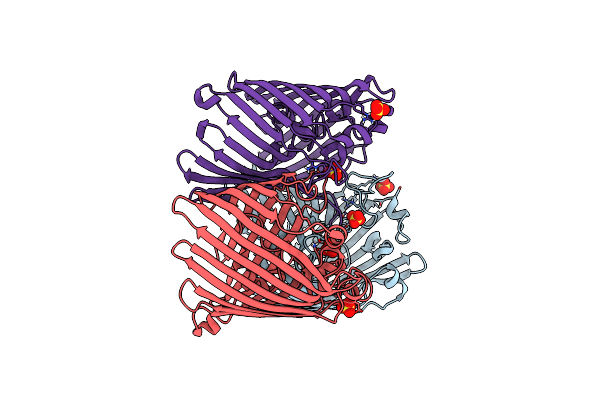 |
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN Ligands: SO4 |
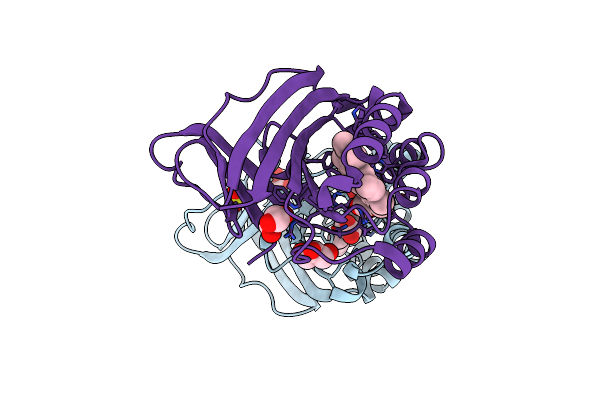 |
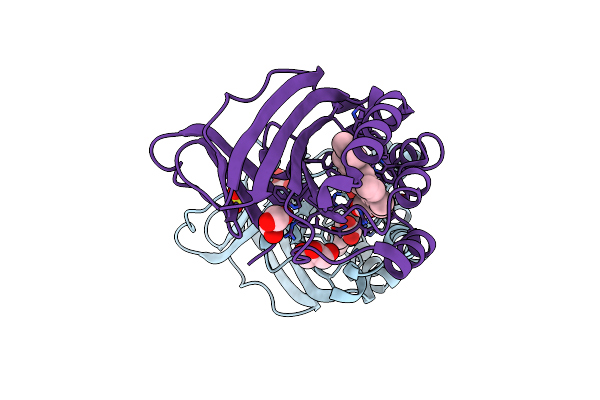
Crystal Structure Of Chlorite Dismutase At 3000 Ev Based On Spherical Harmonics Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-19 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |
 |
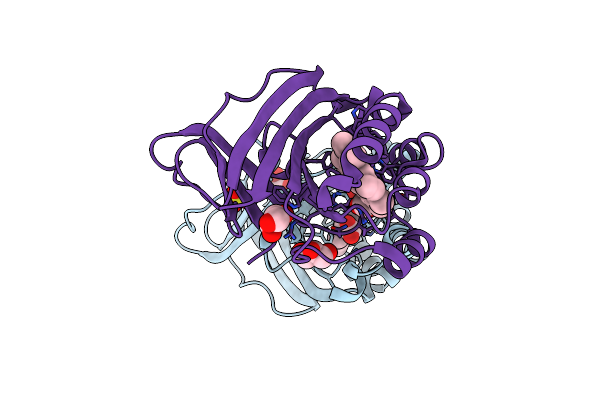
Crystal Structure Of Chlorite Dismutase At 3000 Ev With No Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-19 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |
 |
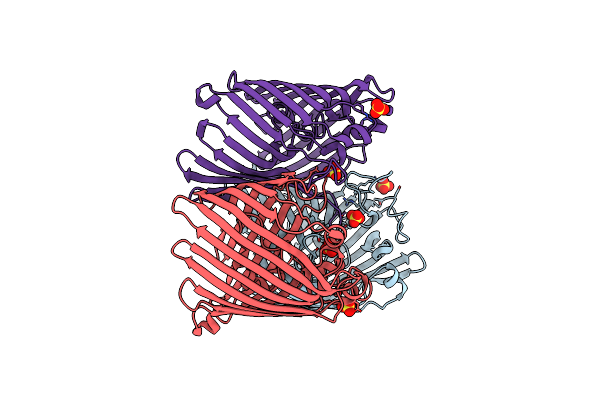
Crystal Structure Of Chlorite Dismutase At 3000 Ev Based On A Combination Of Spherical Harmonics And Analytical Absorption Corrections
Organism: Cyanothece sp. pcc 7425
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2024-06-19 Classification: OXIDOREDUCTASE Ligands: HEM, SO4, GOL, CL |
 |
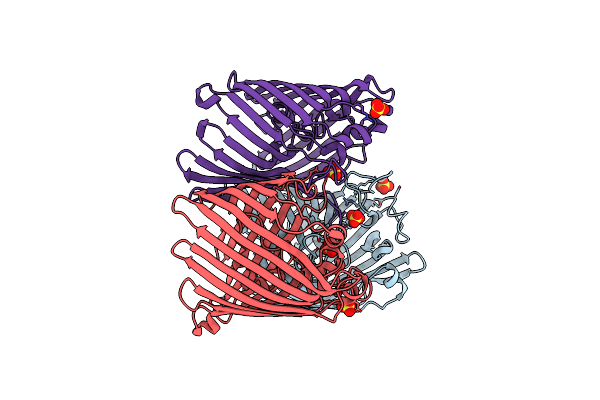
Crystal Structure Of Ompk36 Gd At 3500 Ev Based On A Combination Of Spherical Harmonics And Analytical Absorption Corrections
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN Ligands: SO4 |
 |
Crystal Structure Of Ompk36 Gd At 3500 Ev Based On Analytical Absorption Corrections
Organism: Klebsiella pneumoniae
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2024-06-19 Classification: MEMBRANE PROTEIN Ligands: SO4 |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-02-28 Classification: HYDROLASE Ligands: CL |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-02-28 Classification: HYDROLASE Ligands: CL |
 |
Ssx Structure Of Arabidopsis Thaliana Pdx1.3 Grown In Seeded Batch Conditions
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2024-02-28 Classification: LYASE Ligands: PO4 |

