Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
Planned Maintenance: Some services may turn out to be unavailable from 15th January, 2026 to 16th January, 2026. We apologize for the inconvenience!
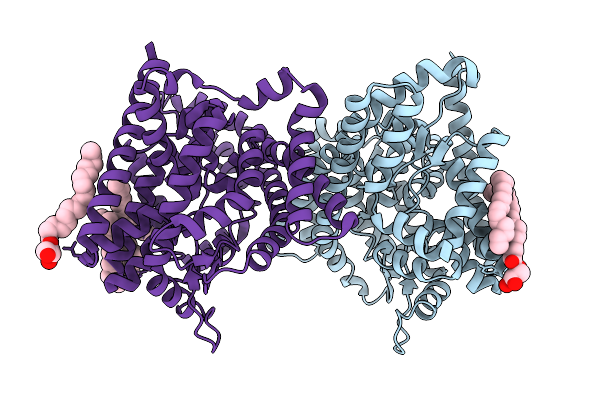 |
Crystal Structure Of Leminorella Grimontii Gatc In The Presence Of D-Xylose
Organism: Leminorella grimontii
Method: X-RAY DIFFRACTION Release Date: 2025-10-15 Classification: TRANSPORT PROTEIN Ligands: OLC |
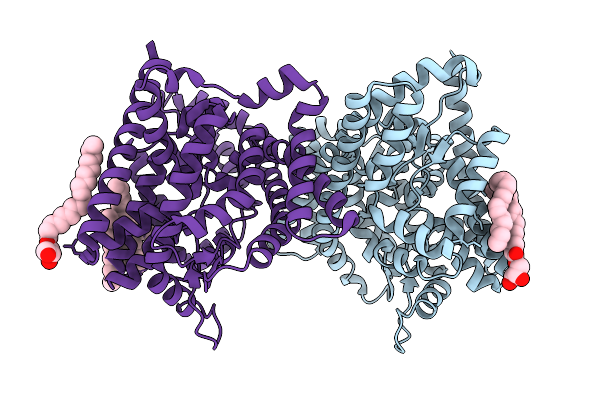 |
Organism: Leminorella grimontii
Method: X-RAY DIFFRACTION Release Date: 2025-10-15 Classification: TRANSPORT PROTEIN Ligands: OLC |
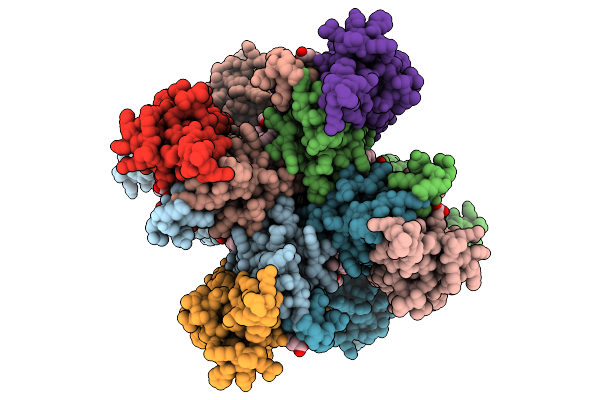 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: PLM, OLC, E2Q |
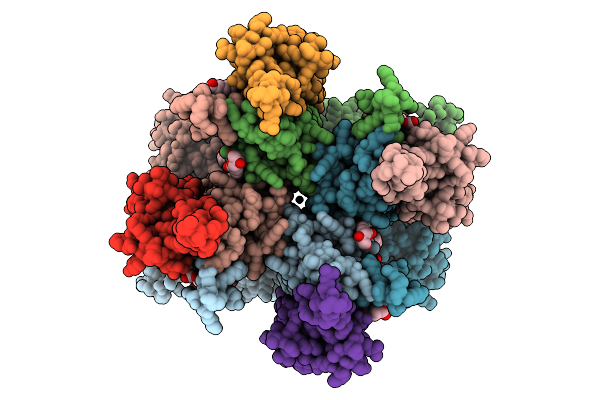 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: PLM, OLC, CYZ, GLU, CA |
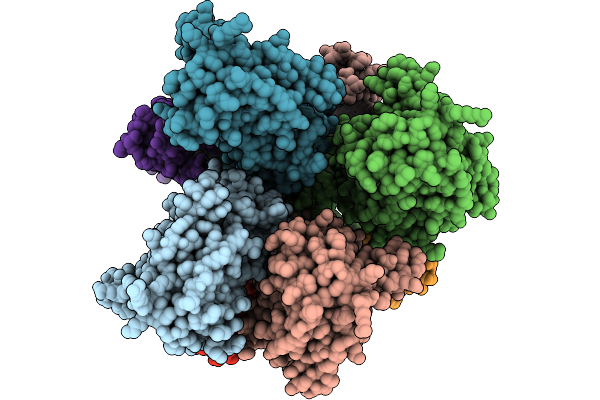 |
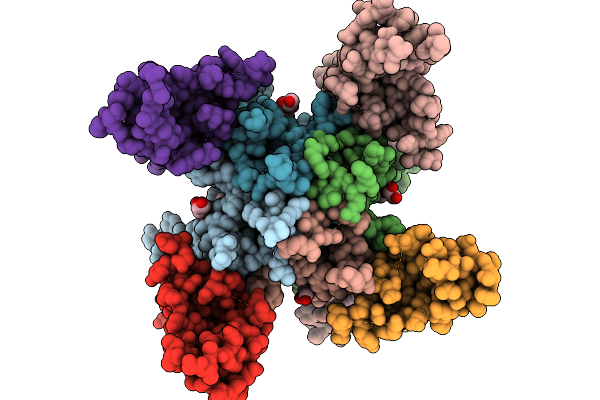
Glua4 In Complex With Tarp-2, Desensitized State, Structure Of Tmd/Lbd Domains
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: PLM, CA, OLC |
 |
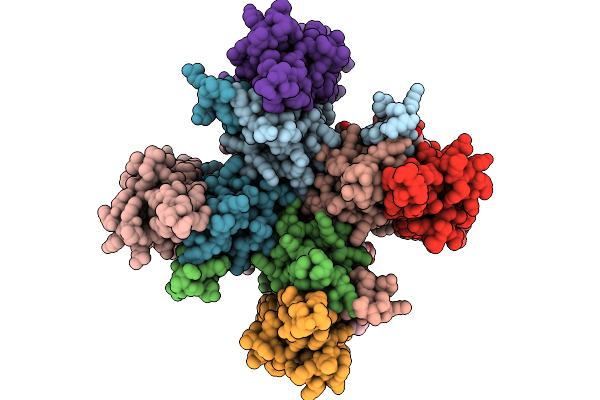
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: PLM, CA, OLC |
 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-24 Classification: MEMBRANE PROTEIN Ligands: PLM, OLC |
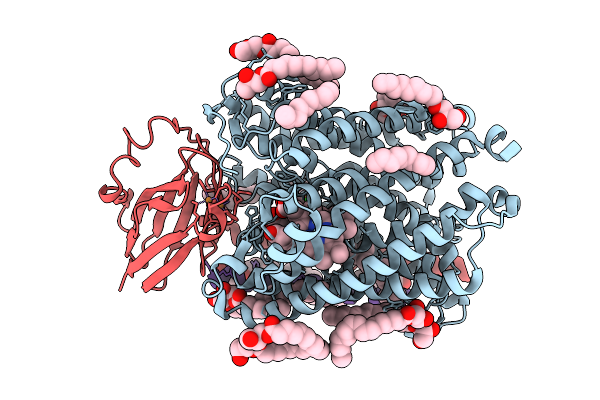 |
Chloride Bound Structure Of Oxidized Ba3-Type Cytochrome C Oxidase Confirmed By Single-Wavelength Anomalous Diffraction
Organism: Thermus thermophilus
Method: X-RAY DIFFRACTION Release Date: 2025-08-13 Classification: OXIDOREDUCTASE Ligands: CU, HEM, HAS, OLC, CL, CUA |
 |
Room Temperature Structure Of Kr2 Rhodopsin In Pentameric Form At 95% Relative Humidity
Organism: Dokdonia eikasta
Method: X-RAY DIFFRACTION Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: RET, NA, OLC, LFA |
 |
Room-Temperature Structure Of Kr2 Rhodopsin In Pentameric Form At 85% Relative Humidity
Organism: Dokdonia eikasta
Method: X-RAY DIFFRACTION Release Date: 2025-07-16 Classification: MEMBRANE PROTEIN Ligands: RET, NA, OLC, LFA |
 |
Cryoem Structure Of Human Full-Length Alpha1Beta3Gamma2 Gaba(A) Receptor In Complex With Garlh4, The Tmd Of Neuroligin2, Gaba And Megabody38, In A Desensitised State (Stated1)
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: MEMBRANE PROTEIN Ligands: PIO, PGW, D10, PX2, ABU, CL, R16, HEX, PLM, OLC, CLR |
 |
Cryoem Structure Of Human Full-Length Alpha1Beta3Gamma2 Gaba(A) Receptor In Complex With Garlh4, The Tmd Of Neuroligin2, Gaba And Megabody38 In A Desensitised State (Stated2)
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: MEMBRANE PROTEIN Ligands: HEX, PIO, PGW, PX2, D10, PLM, CL, ABU, R16, OLC, CLR |
 |
Crystal Structure Of Marine Actinobacteria Clade Rhodopsin (Mar) In The Ground State
Organism: Candidatus actinomarina minuta, Marine actinobacteria clade
Method: X-RAY DIFFRACTION Release Date: 2025-04-02 Classification: MEMBRANE PROTEIN Ligands: OLC, LFA, GOL, RET, PO4 |
 |
Crystal Structure Of Marine Actinobacteria Clade Rhodopsin (Mar) In The P596 State
Organism: Candidatus actinomarina minuta, Marine actinobacteria clade
Method: X-RAY DIFFRACTION Release Date: 2025-04-02 Classification: MEMBRANE PROTEIN Ligands: OLC, LFA, GOL, RET, PO4 |
 |
The Crystal Structure Of Yegtk267A From The Nucleoside: H+ Symporter Family
Organism: Escherichia coli (strain k12)
Method: X-RAY DIFFRACTION Resolution:2.91 Å Release Date: 2025-04-02 Classification: TRANSPORT PROTEIN Ligands: OLC |
 |
Crystal Structure Of The Light-Driven Proton Pump Heimdallarchaeial Rhodopsin Heimdallr1
Organism: Candidatus heimdallarchaeota archaeon
Method: X-RAY DIFFRACTION Release Date: 2025-04-02 Classification: PROTON TRANSPORT Ligands: RET, OLC, NO3 |
 |
Organism: Pseudomonas aeruginosa pao1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2025-03-26 Classification: HYDROLASE Ligands: OLC |
 |
True-Atomic Resolution Crystal Structure Of The Open State Of The Viral Channelrhodopsin Olpvr1
Organism: Organic lake phycodnavirus
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2025-03-12 Classification: MEMBRANE PROTEIN Ligands: RET, NA, 97N, OLC, LFA, GOL |
 |
Structure Of The Viral Channelrhodopsin Olpvr1 At Low Ph Obtained By Soaking
Organism: Organic lake phycodnavirus
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2025-03-12 Classification: MEMBRANE PROTEIN Ligands: RET, 97N, LFA, OLC |
 |
Organism: Dokdonia eikasta
Method: ELECTRON MICROSCOPY Resolution:2.32 Å Release Date: 2025-01-29 Classification: MEMBRANE PROTEIN Ligands: OLC, LFA, RET, NA, OLA |