Search Count: 55
 |
Organism: Bacillota bacterium
Method: ELECTRON MICROSCOPY Resolution:4.10 Å Release Date: 2025-09-03 Classification: PROTEIN FIBRIL |
 |
Crystal Structure Of Complement1.4 (Cent1.4), An Engineered Photoenzyme For Selective [2+2]-Cycloadditions
Organism: Loligo vulgaris
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, PEG, PGE |
 |
C-Terminal Domain Of The F-Ena Tip Fibrillum F-Bcla From Bacillus Thuringiensis
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2025-03-12 Classification: STRUCTURAL PROTEIN Ligands: GOL, EDO |
 |
F-Ena Exosporium Anchoring Complex Between Exsf And A Peptide Derived From The N-Terminus Of F-Anchor
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2025-03-12 Classification: STRUCTURAL PROTEIN |
 |
Cryoem Structure Of F-Ena Fibers On The Spores Of Bacillus Thuringiensis Serovar Kurstaki
Organism: Bacillus thuringiensis serovar kurstaki
Method: ELECTRON MICROSCOPY Release Date: 2025-03-12 Classification: PROTEIN FIBRIL |
 |
Organism: Bacillus thuringiensis serovar kurstaki
Method: ELECTRON MICROSCOPY Release Date: 2025-03-12 Classification: PROTEIN FIBRIL |
 |
Organism: Cohnella sp. ov330
Method: ELECTRON MICROSCOPY Resolution:4.10 Å Release Date: 2025-03-12 Classification: PROTEIN FIBRIL |
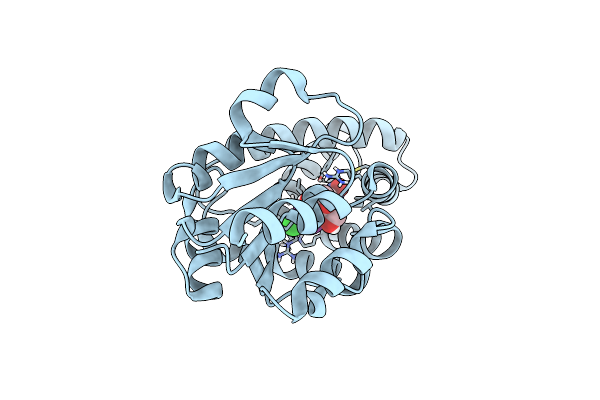 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2025-01-29 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, A1IGD, CL |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2025-01-29 Classification: BIOSYNTHETIC PROTEIN Ligands: IOD, PG4 |
 |
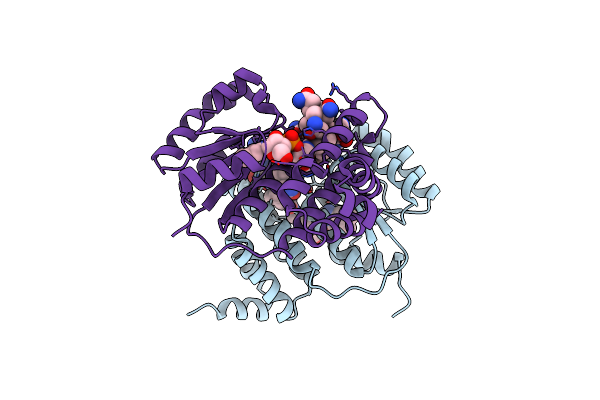
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2025-01-29 Classification: BIOSYNTHETIC PROTEIN Ligands: PG4, PEG, CL, PGE |
 |
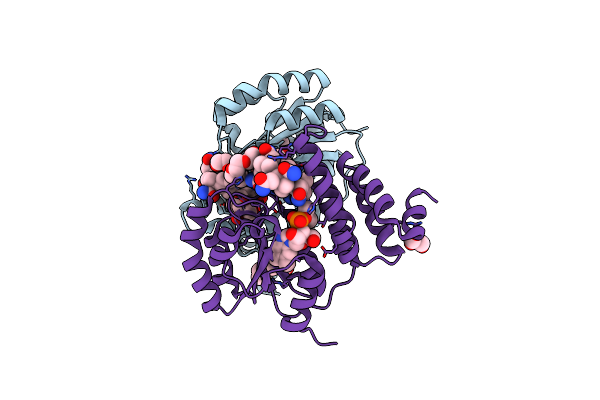
B12-Binding Domain From Chloracidobacterium Thermophilum Merr Family Protein, Dark State
Organism: Chloracidobacterium thermophilum
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-11-13 Classification: TRANSCRIPTION Ligands: B12, 5AD |
 |
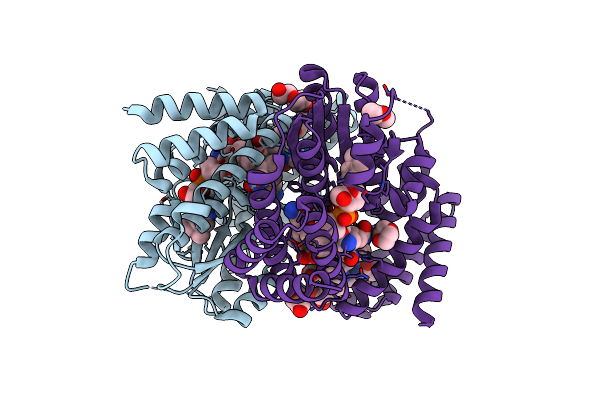
B12-Binding Domain From Chloracidobacterium Thermophilum Merr Family Protein, Anaerobic Light State
Organism: Chloracidobacterium thermophilum
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-11-13 Classification: TRANSCRIPTION Ligands: B12, TRS, P6G, PG4, GOL |
 |
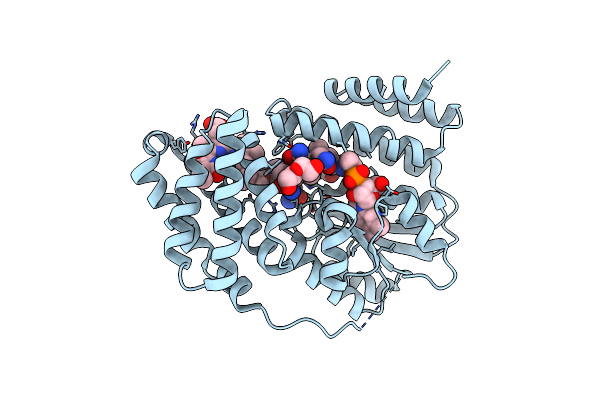
Organism: Saccharothrix syringae
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-04-10 Classification: UNKNOWN FUNCTION Ligands: B12, 5AD, BLA, DIO, PEG |
 |
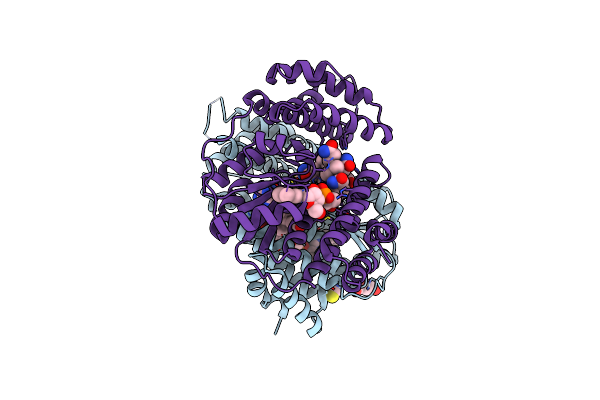
Organism: Saccharothrix syringae
Method: X-RAY DIFFRACTION Resolution:1.98 Å Release Date: 2024-04-10 Classification: UNKNOWN FUNCTION Ligands: B12, BLA, PEG |
 |
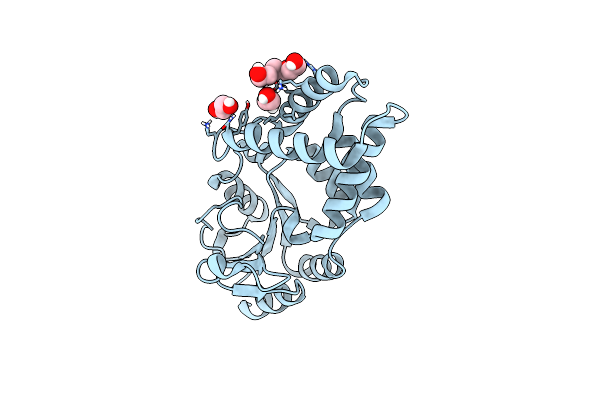
Organism: Acidimicrobiaceae bacterium
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2024-04-10 Classification: UNKNOWN FUNCTION Ligands: B12, 5AD, PEG, SCN |
 |
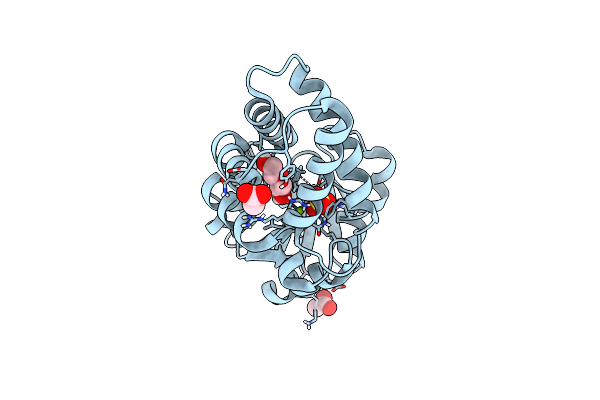
Crystal Structure Of Bhmehis1.8, An Engineered Enzyme For The Morita-Baylis-Hillman Reaction
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2024-02-28 Classification: BIOSYNTHETIC PROTEIN Ligands: PGE, EDO |
 |
Crystal Structure Of Bhmehis1.0, An Engineered Enzyme For The Morita-Baylis-Hillman Reaction
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2024-02-28 Classification: BIOSYNTHETIC PROTEIN Ligands: PEG, ACT, MG, SO4 |
 |
Dark State 1.8 Angstrom Crystal Structure Of Cobalamin Binding Domain Belonging To A Light-Dependent Transcription Regulator Ttcarh Obtained Under Aerobic Condition
Organism: Thermus thermophilus hb27
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2023-08-16 Classification: TRANSCRIPTION Ligands: BR, B12, 5AD, IMD, PGE, EDO |
 |
Dark State 2.2 Angstrom Crystal Structure Of Cobalamin Binding Domain Belonging To A Light-Dependent Transcription Regulator Ttcarh Obtained Under Anaerobic Condition
Organism: Thermus thermophilus hb27
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2023-08-16 Classification: TRANSCRIPTION Ligands: BR, B12, 5AD |
 |
Anaerobic Light Exposed 2.25 Angstrom Crystal Structure Of Cobalamin Binding Domain Belonging To A Light-Dependent Transcription Regulator Ttcarh
Organism: Thermus thermophilus hb27
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2023-08-16 Classification: TRANSCRIPTION Ligands: B12, BR |

