Search Count: 8
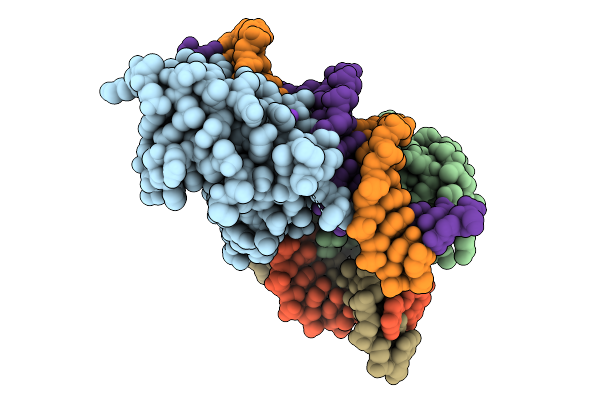 |
Co-Crystal Structure Of Yeast Forkhead Transcription Factor Fkh1 Bound To Dna
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2025-09-24 Classification: DNA BINDING PROTEIN/DNA Ligands: K |
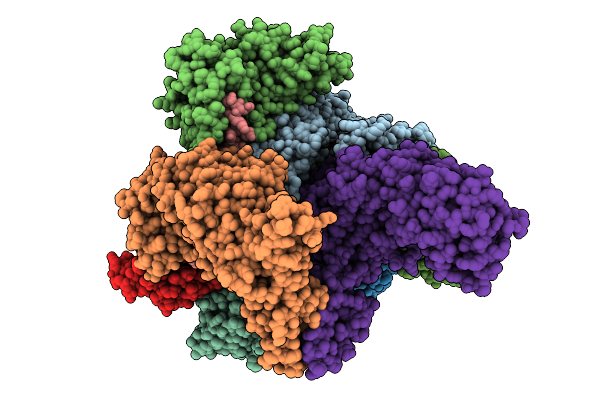 |
Structure Of Spin90 Dimer-Arp2/3 Complex-Nucleated Actin Filament (Singlet Complex)
Organism: Homo sapiens, Bos taurus, Gallus gallus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: CYTOSOLIC PROTEIN Ligands: MG, ADP |
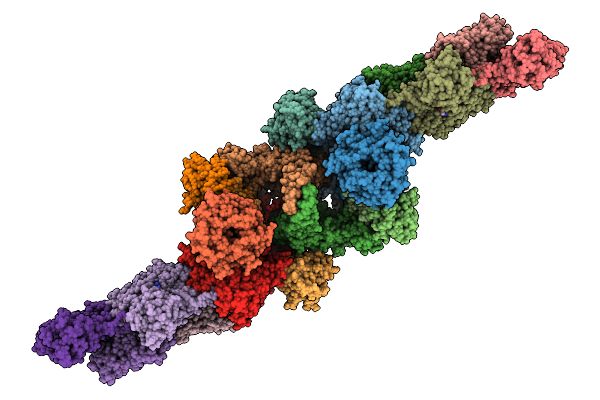 |
Structure Of Spin90 Dimer-Arp2/3 Complexes-Nucleated Actin Filaments (Doublet Complex)
Organism: Homo sapiens, Bos taurus, Gallus gallus
Method: ELECTRON MICROSCOPY Release Date: 2025-09-10 Classification: CYTOSOLIC PROTEIN Ligands: ADP, MG |
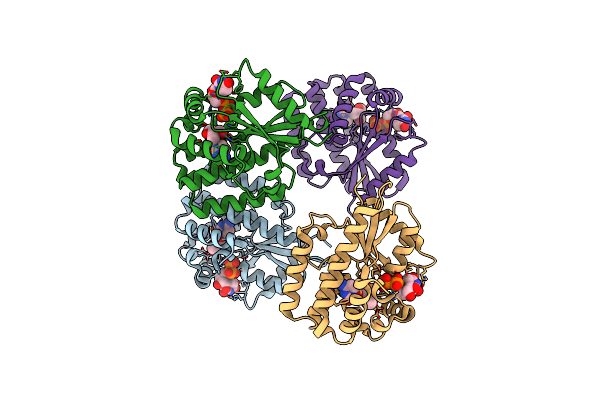 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-12-20 Classification: TRANSFERASE Ligands: UDP, VVC |
 |
The Structure Of Nica2 Variant F104L/A107T/S146I/G317D/H368R/L449V/N462S From Pseudomonas Putida
Organism: Pseudomonas putida s16
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2023-07-26 Classification: FLAVOPROTEIN Ligands: FDA |
 |
The Structure Of Nica2 Variant F104L/A107T/S146I/G317D/H368R/L449V/N462S In Complex With N-Methylmyosmine
Organism: Pseudomonas putida s16
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2023-07-26 Classification: FLAVOPROTEIN Ligands: NCT, FDA, EDO |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2023-07-26 Classification: FLAVOPROTEIN, OXIDOREDUCTASE Ligands: FDA, PG4, PEG, TXU |
 |
Organism: Ideonella sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2020-09-23 Classification: HYDROLASE Ligands: SO4 |

