Search Count: 17
All
Selected
 |
Low-Dose (14.9 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Room Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Higher-Dose (283.1 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Room Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Low-Dose (33.3 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Cryogenic Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Higher-Dose (1332 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Cryogenic Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.24 Å Release Date: 2026-03-04 Classification: OXYGEN STORAGE Ligands: HEM |
 |
Higher-Dose (260.0 Kgy) Structure Of Horse-Heart Myoglobin At Room Temperature
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Higher-Dose (676.8 Kgy) Structure Of Horse-Heart Myoglobin At Cryogenic Temperature
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2026-03-04 Classification: OXYGEN STORAGE Ligands: HEM, MLA, NA |
 |
Low-Dose (14.4 Kgy) Structure Of Horse-Heart Myoglobin At Cryogenic Temperature
Organism: Equus caballus
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2026-03-04 Classification: OXYGEN STORAGE Ligands: HEM, MLI, NA |
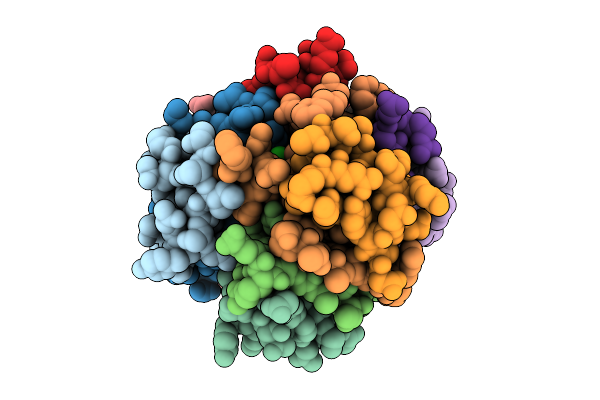 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-10-29 Classification: HORMONE Ligands: CRS, ZN, CL, CA |
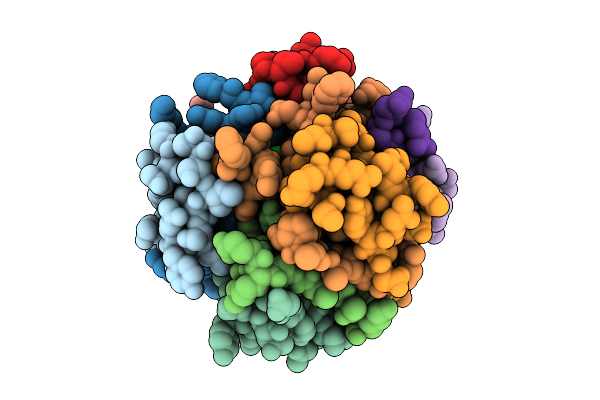 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.38 Å Release Date: 2025-10-29 Classification: HORMONE Ligands: IPH, ZN, CL, CA |
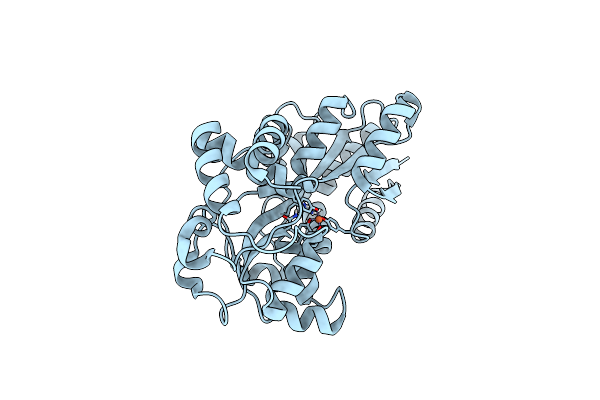 |
Organism: Prochlorococcus marinus subsp. pastoris str. ccmp1986
Method: NEUTRON DIFFRACTION Resolution:2.10 Å Release Date: 2024-01-17 Classification: METAL BINDING PROTEIN Ligands: FE |
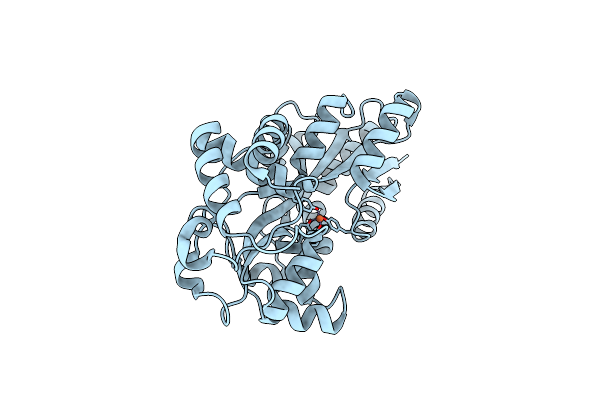 |
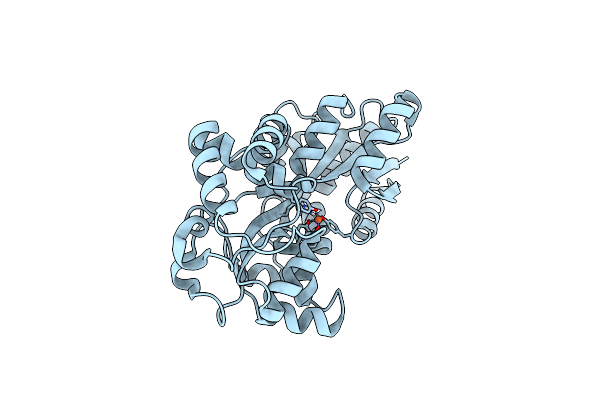
Organism: Prochlorococcus marinus subsp. pastoris str. ccmp1986
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2023-08-30 Classification: METAL BINDING PROTEIN Ligands: FE |
 |
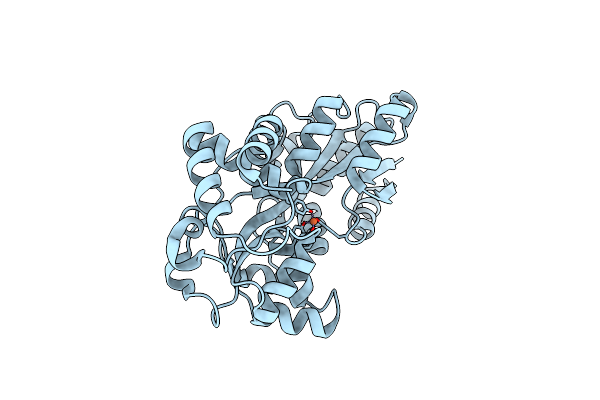
Organism: Prochlorococcus marinus subsp. pastoris str. ccmp1986
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2023-08-30 Classification: METAL BINDING PROTEIN Ligands: FE |
 |
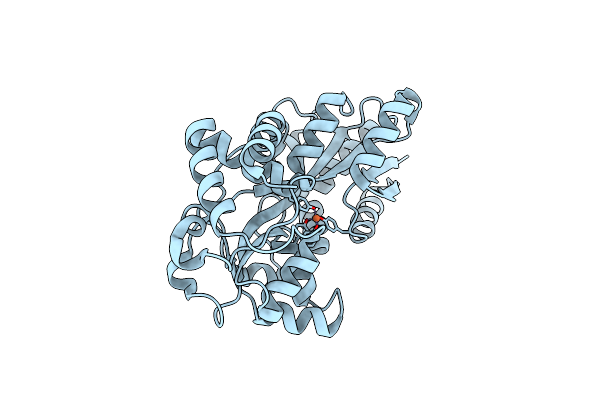
Organism: Prochlorococcus marinus subsp. pastoris str. ccmp1986
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2023-08-30 Classification: METAL BINDING PROTEIN Ligands: FE2 |
 |
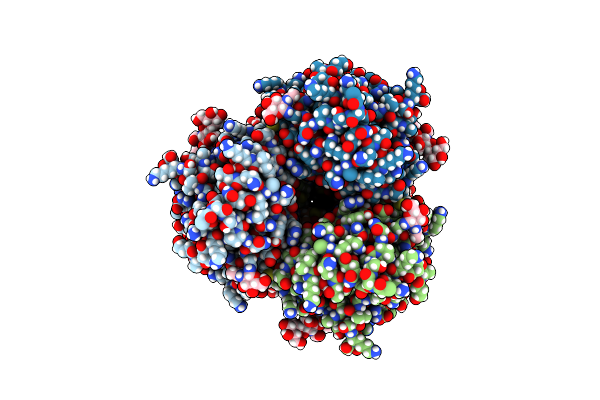
Organism: Prochlorococcus marinus subsp. pastoris str. ccmp1986
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2023-08-30 Classification: METAL BINDING PROTEIN Ligands: FE |
 |
Cryo-Em Structure Of Designed Influenza Ha Binder, Ha_20, Bound To Influenza Ha (Strain: Iowa43)
Organism: Influenza a virus, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2023-06-14 Classification: DE NOVO PROTEIN/Viral Protein Ligands: NAG |
 |
Elongated Version Of A De Novo Designed Three Helix Bundle Structure (Gra3D)
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2018-10-03 Classification: DE NOVO PROTEIN Ligands: ZN, SCN, CL, EDO |

