Search Count: 3,467
All
Selected
 |
Structure Of The Lactate Monooxygenase Mutant Y44N, Y152F, H290S, T181A, F184P
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2026-02-18 Classification: OXIDOREDUCTASE Ligands: FMN, GOL, CIT, SO4 |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD, CL |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis (Alternative Loop Conformation)
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis (Alternative N-Terminus)
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FE |
 |
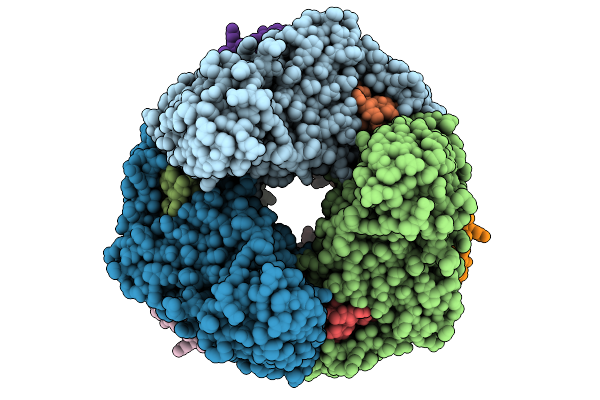
Organism: Nitrosomonas halophila
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: MEMBRANE PROTEIN Ligands: A1EZJ, DKA, PLM, ZP7, CU, A1EZI, 4AG |
 |
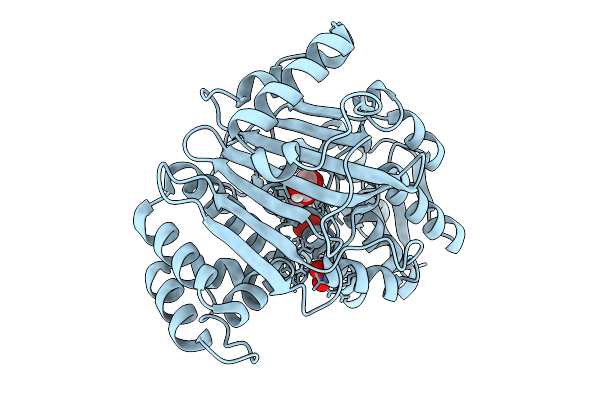
Organism: Nitrosospira briensis
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: OXIDOREDUCTASE Ligands: CU, 6PL |
 |
Organism: Rhodococcus
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-01-28 Classification: OXIDOREDUCTASE Ligands: NO3, PG0 |
 |
Organism: Pseudomonas putida
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, ACT |
 |
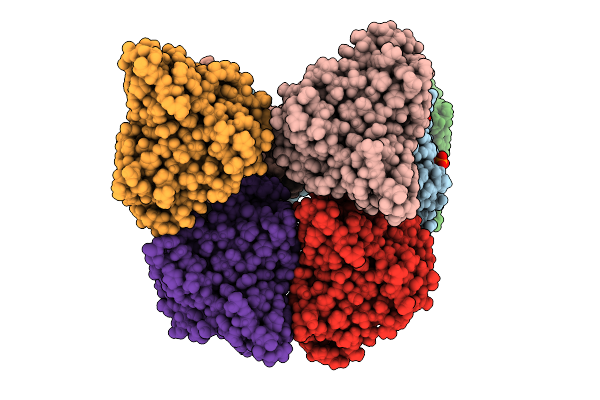
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-01-07 Classification: OXIDOREDUCTASE Ligands: HEM, A1EI5 |
 |
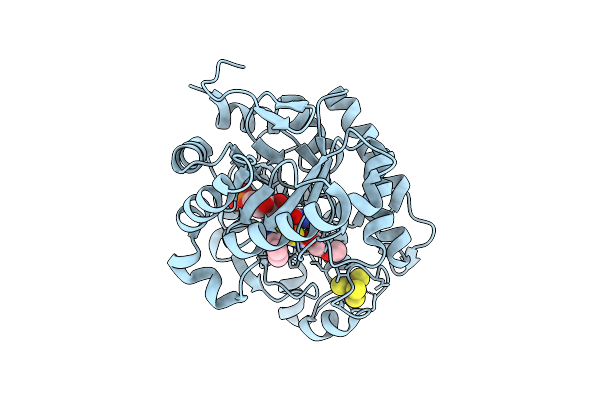
Organism: Cellvibrio japonicus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-12-24 Classification: OXIDOREDUCTASE Ligands: CU, PO4 |
 |
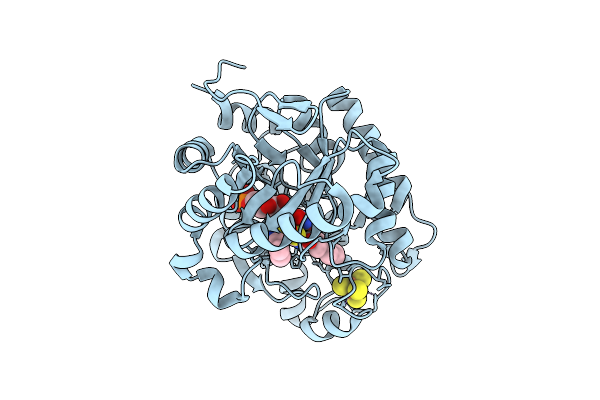
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 1
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: SF4, A1ELE, FMN |
 |
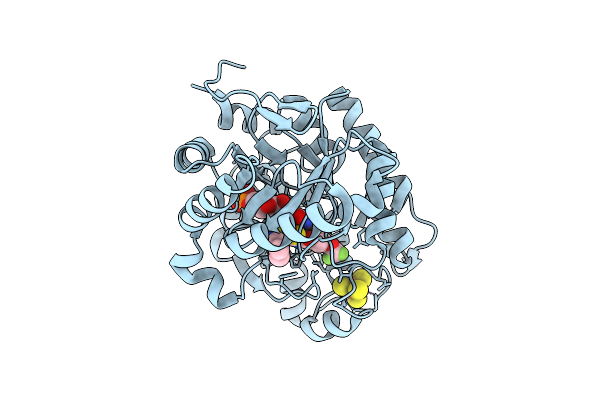
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 21
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELF |
 |
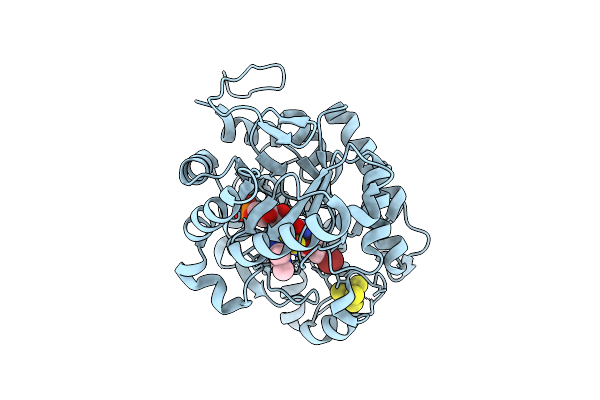
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 11
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELG |
 |
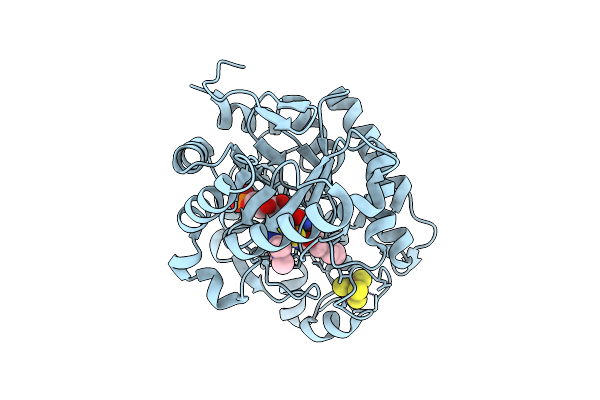
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 35
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1ELH |
 |
Crystal Structure Of Dehydrogenase/Isomerase Fabx From Helicobacter Pylori In Complex With Inhibitor 47
Organism: Helicobacter pylori
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2025-12-24 Classification: BIOSYNTHETIC PROTEIN Ligands: FMN, SF4, A1EMD |
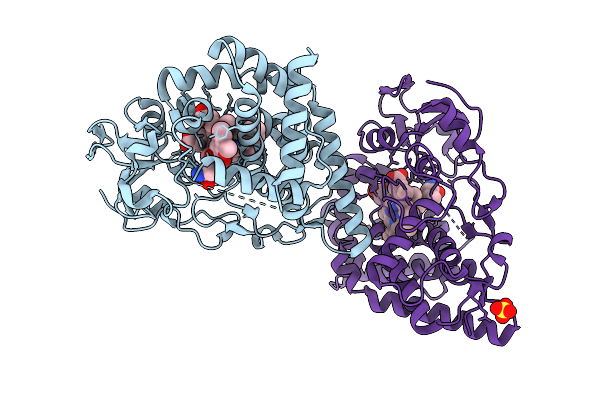 |
Organism: Streptomyces peucetius subsp. caesius atcc 27952
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2025-12-24 Classification: OXIDOREDUCTASE Ligands: HEM, DM1, SO4 |
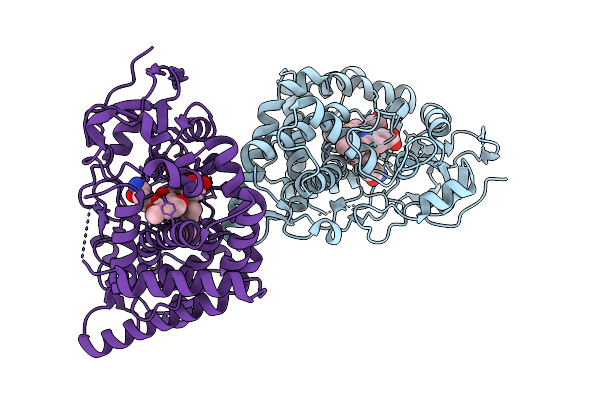 |
Organism: Streptomyces peucetius
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: HEM, A1L3V |
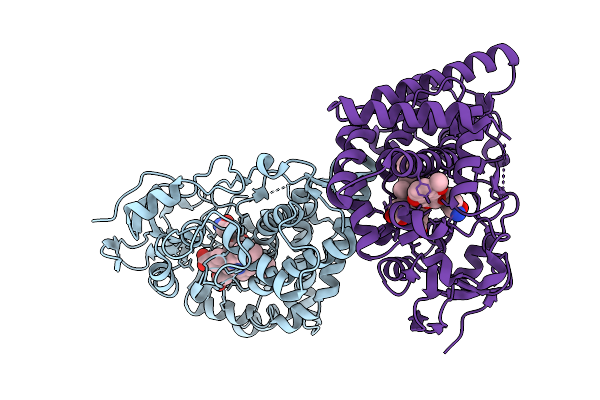 |
Organism: Streptomyces peucetius
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2025-12-24 Classification: TRANSFERASE Ligands: HEM, A1JN3 |
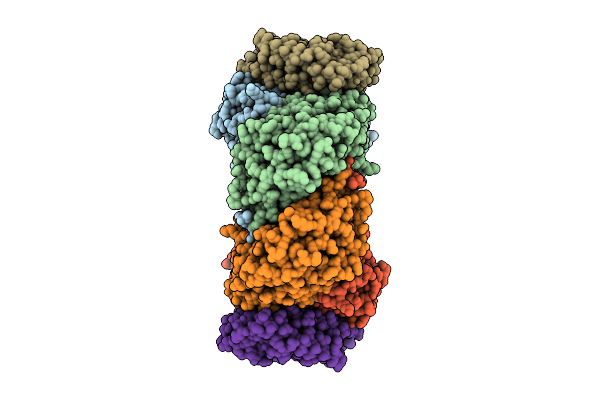 |
Cryo-Em Structure Of Soluble Methane Monooxygenase Hydroxylase From Methylosinus Sporium 5
Organism: Methylosinus sporium
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: OXIDOREDUCTASE Ligands: FE |

