Search Count: 478
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: A1EY6, EDO |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: GOL, A1EY5 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: A1EY4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: GOL, EDO, A1EY3 |
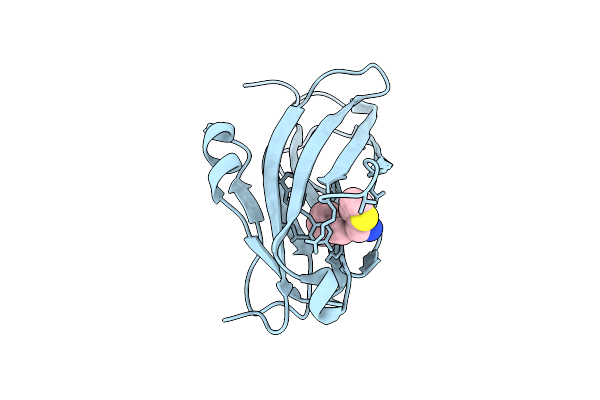 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Release Date: 2025-10-29 Classification: SIGNALING PROTEIN Ligands: A1EY2 |
 |
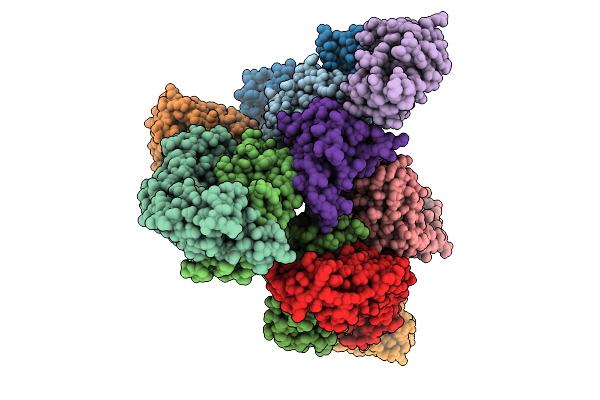
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-01 Classification: EXOCYTOSIS Ligands: GNP |
 |
Organism: Mus musculus, Monkeypox virus
Method: X-RAY DIFFRACTION Release Date: 2025-10-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2025-10-01 Classification: METAL BINDING PROTEIN |
 |
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2025-10-01 Classification: METAL BINDING PROTEIN |
 |
Solution Structure Of Silver Bound Xpc Binding Domain Of Hhr23B (Holo Form)
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2025-10-01 Classification: METAL BINDING PROTEIN Ligands: AG |
 |
Organism: Mus musculus, Monkeypox virus
Method: X-RAY DIFFRACTION Release Date: 2025-10-01 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-08-06 Classification: MEMBRANE PROTEIN |
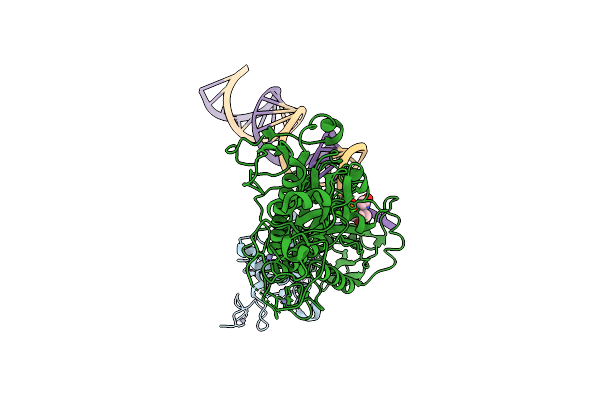 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: VIRAL PROTEIN/RNA Ligands: ZN, CA, K5X |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2025-07-16 Classification: VIRAL PROTEIN/RNA Ligands: ZN, CA, EIF |
 |
Organism: Felis catus
Method: X-RAY DIFFRACTION Release Date: 2025-07-02 Classification: IMMUNE SYSTEM |
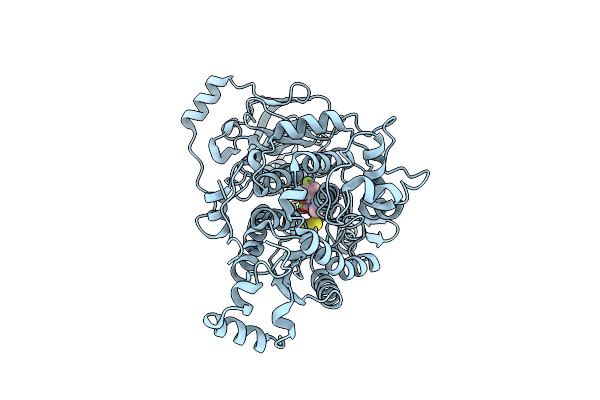 |
Organism: Arabidopsis thaliana
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2025-06-04 Classification: LYASE Ligands: FES, A1L7A, MG |
 |
Mycobacterium Tuberculosis Pks13 Acyltransferase Serine Converted To Beta-Lactam Form By Cec215 Via Sufex Reaction
Organism: Mycobacterium tuberculosis h37rv
Method: X-RAY DIFFRACTION Release Date: 2025-05-21 Classification: ANTIBIOTIC Ligands: SO4, CL, PG4, EDO, DMS, GOL, PEG |
 |
Organism: Mycobacterium tuberculosis (strain atcc 25618 / h37rv)
Method: X-RAY DIFFRACTION Release Date: 2025-05-21 Classification: ANTIBIOTIC Ligands: DMS, SO4, PE5, EDO, PEG, GOL, CL, NA, P33 |

