Search Count: 222
All
Selected
 |
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: A1JCI |
 |
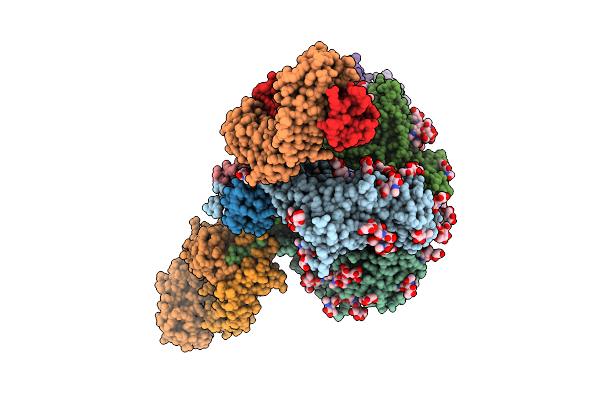
The Structure Of Mcat1 In Complex With Its Substrate Ornithine And The Rbd Of Frmlv.
Organism: Mus musculus, Friend murine leukemia virus (isolate 57)
Method: ELECTRON MICROSCOPY Release Date: 2026-02-11 Classification: MEMBRANE PROTEIN Ligands: ORN, NAG |
 |
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1IXC |
 |
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1IXB, PO4 |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: CELL ADHESION |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: CELL ADHESION |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: CELL ADHESION |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: CELL ADHESION |
 |
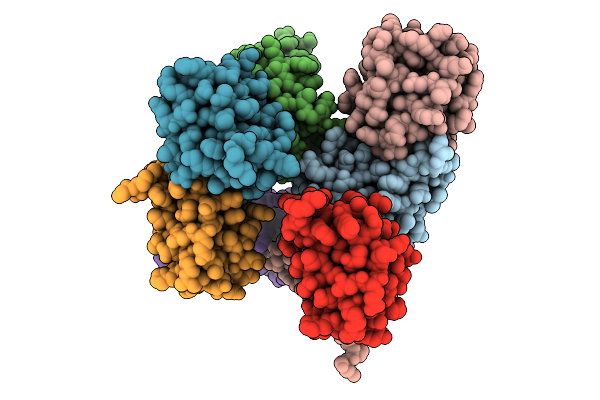
Cryo-Em Structure Of Hiv-1 Bg505Ds-Sosip.664 Env Trimer Bound To Dfph-A.01_10R59P_Lc Fab
Organism: Human immunodeficiency virus 1, Macaca
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: VIRAL PROTEIN Ligands: NAG |
 |
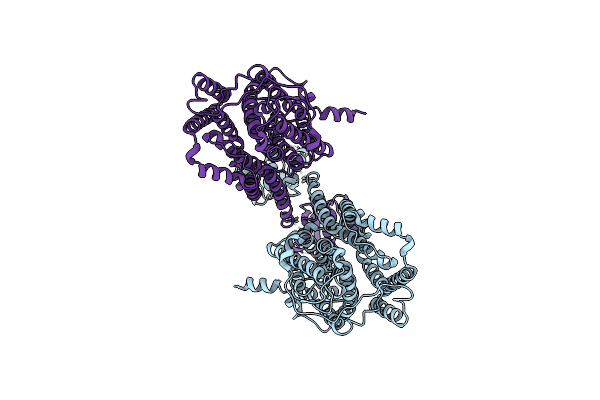
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2025-10-08 Classification: SIGNALING PROTEIN |
 |
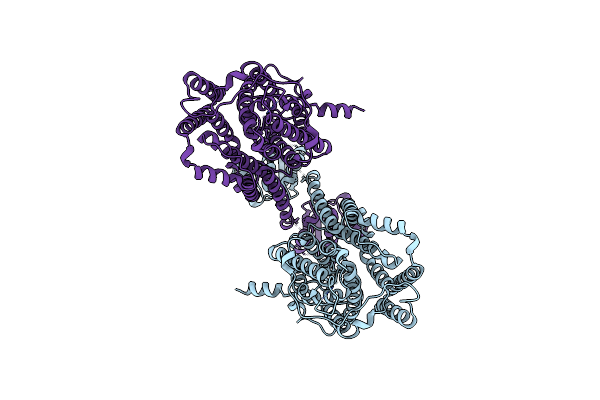
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: TRANSPORT PROTEIN Ligands: IOD |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-02 Classification: TRANSPORT PROTEIN |
 |
Organism: Homo sapiens, Vicugna pacos
Method: ELECTRON MICROSCOPY Resolution:3.14 Å Release Date: 2025-06-11 Classification: MEMBRANE PROTEIN Ligands: UTP |
 |
Organism: Homo sapiens, Vicugna pacos
Method: ELECTRON MICROSCOPY Resolution:2.83 Å Release Date: 2025-06-11 Classification: MEMBRANE PROTEIN Ligands: ATP |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2025-06-11 Classification: MEMBRANE PROTEIN Ligands: ATP |
 |
Organism: Homo sapiens, Vicugna pacos
Method: ELECTRON MICROSCOPY Resolution:3.31 Å Release Date: 2025-06-11 Classification: MEMBRANE PROTEIN |
 |
Human Ppar Alpha Ligand Binding Domain In Complex With A 1H-Pyrazolo[3,4-B]Pyridine-Derived Compound
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2025-04-09 Classification: TRANSCRIPTION Ligands: A1LZ9 |
 |
Human Ppar Alpha Ligand Binding Domain In Complex With A 1H-Pyrazolo[3,4-B]Pyridine-Derived Compound
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2025-04-09 Classification: TRANSCRIPTION Ligands: A1L0A |
 |
Human Ppar Alpha Ligand Binding Domain In Complex With A 1H-Pyrazolo[3,4-B]Pyridine-Derived Compound
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-04-02 Classification: TRANSCRIPTION Ligands: A1LZX |
 |
Human Ppar Alpha Ligand Binding Domain In Complex With A 1H-Pyrazolo[3,4-B]Pyridine-Derived Compound
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2025-04-02 Classification: TRANSCRIPTION Ligands: A1LZY |

