Search Count: 21
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Release Date: 2025-10-29 Classification: DE NOVO PROTEIN Ligands: ZN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Release Date: 2025-10-22 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2025-02-19 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.21 Å Release Date: 2025-02-19 Classification: HYDROLASE |
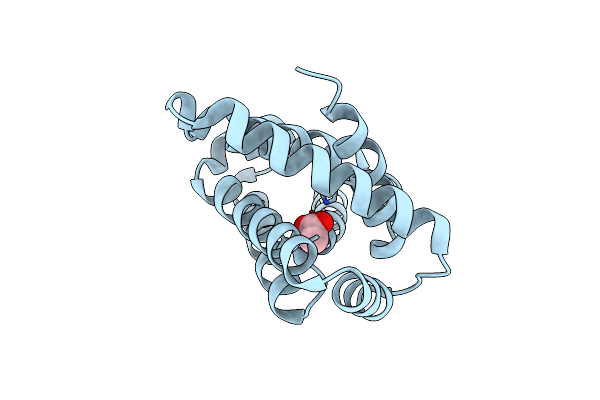 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2025-02-19 Classification: HYDROLASE Ligands: ACY, CL |
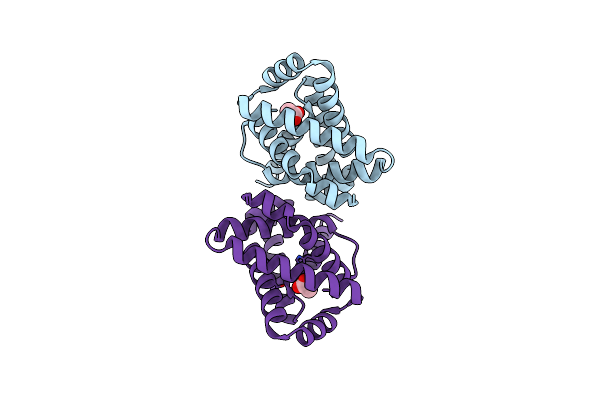 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2025-02-19 Classification: HYDROLASE Ligands: ACT |
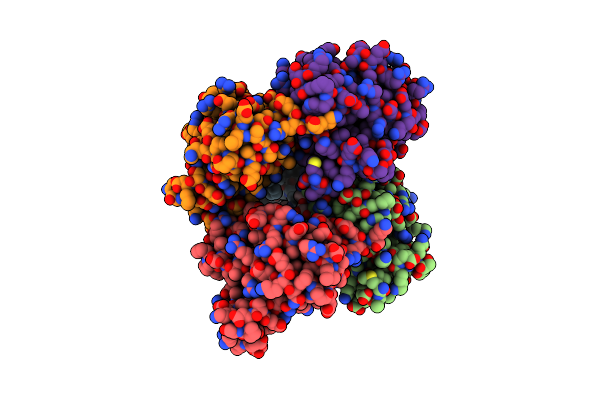 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.21 Å Release Date: 2025-02-19 Classification: HYDROLASE Ligands: PG4, TLA |
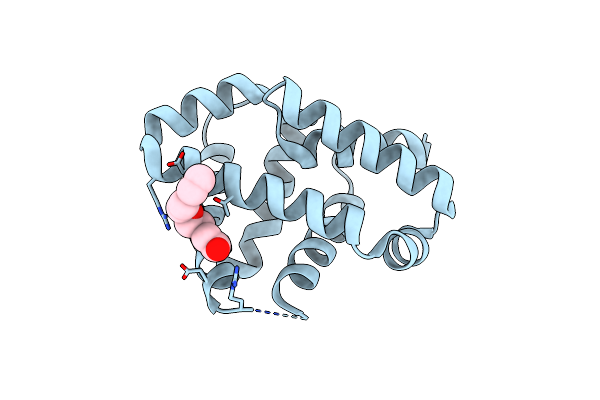 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2025-02-19 Classification: DE NOVO PROTEIN Ligands: PG4 |
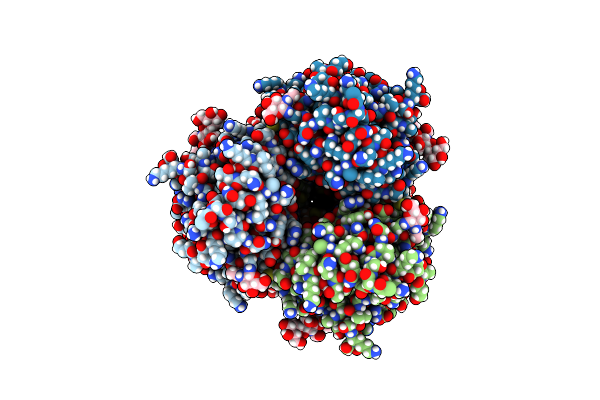 |
Cryo-Em Structure Of Designed Influenza Ha Binder, Ha_20, Bound To Influenza Ha (Strain: Iowa43)
Organism: Influenza a virus, Unidentified
Method: ELECTRON MICROSCOPY Release Date: 2023-06-14 Classification: DE NOVO PROTEIN/Viral Protein Ligands: NAG |
 |
Solution Nmr Structure Of 9-Residue Rosetta-Designed Cyclic Peptide D9.16 In D6-Dmso With Cis/Trans Switching
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
 |
Solution Nmr Structure Of 8-Residue Rosetta-Designed Cyclic Peptide D8.31 In D6-Dmso With Cis/Trans Switching (A-Cc Conformation)
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
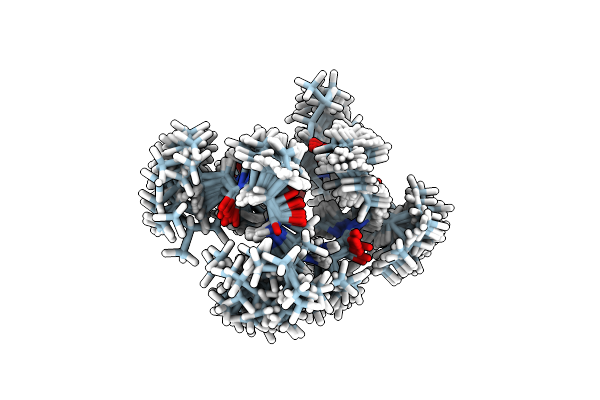 |
Solution Nmr Structure Of 8-Residue Rosetta-Designed Cyclic Peptide D8.21 In D6-Dmso With Cis/Trans Switching
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
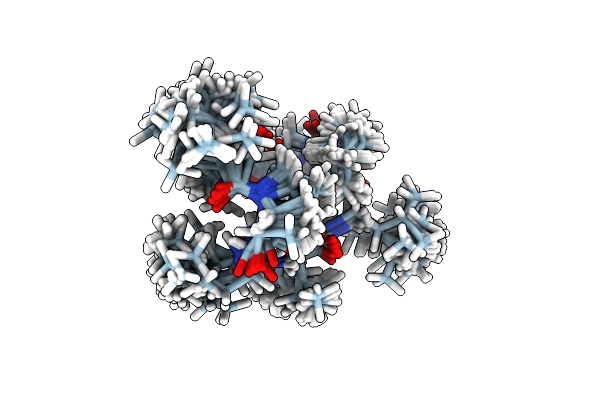 |
Solution Nmr Structure Of 8-Residue Rosetta-Designed Cyclic Peptide D8.21 In 50% D6-Dmso And 50% Water With Cis/Trans Switching (Cc Conformation, 50%)
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
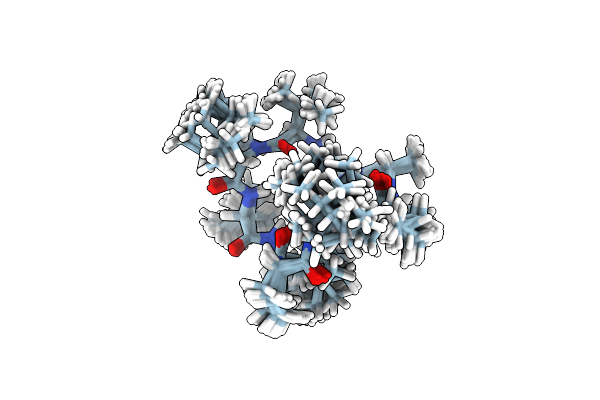 |
Solution Nmr Structure Of 9-Residue Rosetta-Designed Cyclic Peptide D9.16 In Cdcl3 With Cis/Trans Switching (A-Tt Conformation)
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
 |
Solution Nmr Structure Of 8-Residue Rosetta-Designed Cyclic Peptide D8.31 In Cdcl3 With Cis/Trans Switching
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
 |
Solution Nmr Structure Of 8-Residue Rosetta-Designed Cyclic Peptide D8.21 In Cdcl3 With Cis/Trans Switching (Tt Conformation, 47%)
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:0.85 Å Release Date: 2022-09-14 Classification: DE NOVO PROTEIN Ligands: HOH |
 |
Solution Nmr Structure Of 9-Residue Rosetta-Designed Cyclic Peptide D9.16 In Cdcl3 With Cis/Trans Switching (B-Tc Conformation)
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
 |
Solution Nmr Structure Of 8-Residue Rosetta-Designed Cyclic Peptide D8.31 In D6-Dmso With Cis/Trans Switching (B-Ct Conformation)
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |
 |
Solution Nmr Structure Of 8-Residue Rosetta-Designed Cyclic Peptide D8.21 In 50% D6-Dmso And 50% Water With Cis/Trans Switching (Cc Conformation, 50%)
Organism: Synthetic construct
Method: SOLUTION NMR Release Date: 2022-09-14 Classification: DE NOVO PROTEIN |

