Search Count: 19
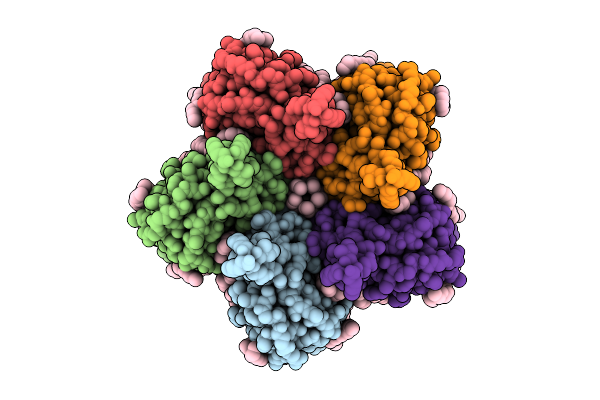 |
Cryo-Em Structure Of The Flotillin-Associated Rhodopsin Psfar In Detergent Micelle
Organism: Candidatus pseudothioglobus
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: MEMBRANE PROTEIN Ligands: LFA |
 |
Cryo-Em Structure Of The Light-Driven Proton Pump Pspr In Detergent Micelle
Organism: Candidatus pseudothioglobus sp.
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: MEMBRANE PROTEIN Ligands: LFA, RET, LMT |
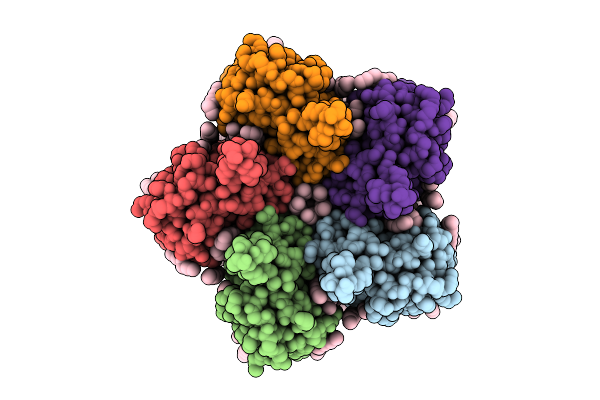 |
Cryo-Em Structure Of The Double Mutant H84V/E120G Of The Flotillin-Associated Rhodopsin Psfar In Detergent Micelle
Organism: Candidatus pseudothioglobus sp.
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: MEMBRANE PROTEIN Ligands: LFA, RET |
 |
Organism: Cryobacterium levicorallinum
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: LFA, RET |
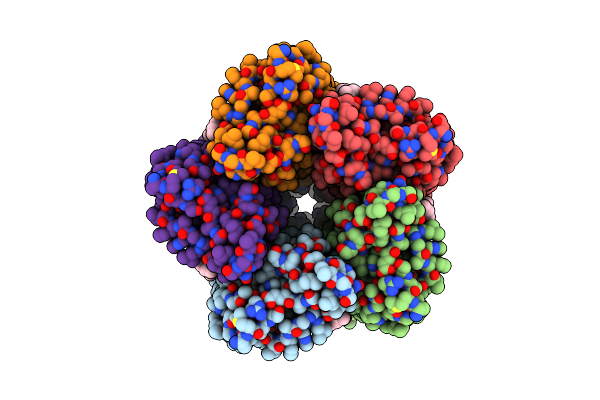 |
Organism: Cryobacterium levicorallinum
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: LFA, RET |
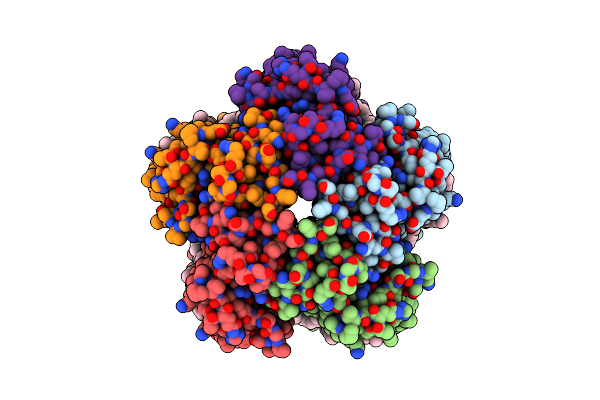 |
Organism: Cryobacterium levicorallinum
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: LMT, LFA, RET |
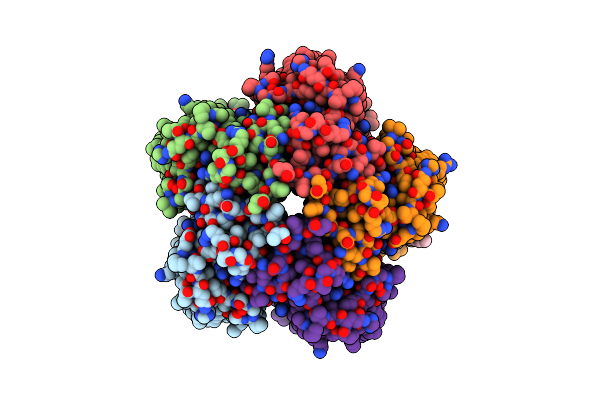 |
Cryo-Em Structure Of The Microbial Rhodopsin Cryor1 At Ph 10.5 In Detergent In The Ground State
Organism: Cryobacterium levicorallinum
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: LMT, RET, LFA |
 |
Cryo-Em Structure Of The Microbial Rhodopsin Cryor1 At Ph 10.5 In Detergent In The M State
Organism: Cryobacterium levicorallinum
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: LMT, RET |
 |
Organism: Subtercola endophyticus
Method: ELECTRON MICROSCOPY Release Date: 2025-05-14 Classification: MEMBRANE PROTEIN Ligands: LFA, RET |
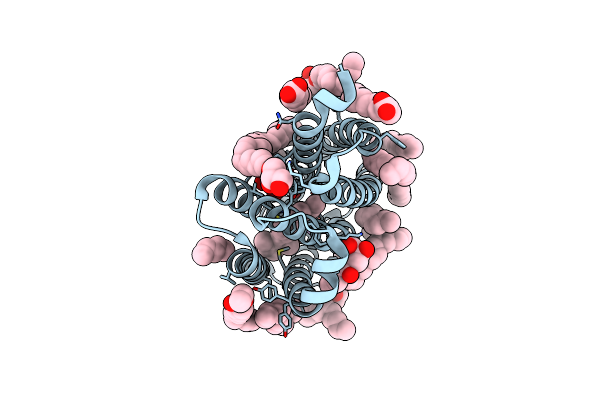 |
True-Atomic Resolution Crystal Structure Of The Closed State Of The Viral Channelrhodopsin Olpvr1
Organism: Organic lake phycodnavirus
Method: X-RAY DIFFRACTION Resolution:1.13 Å Release Date: 2025-03-12 Classification: MEMBRANE PROTEIN |
 |
True-Atomic Resolution Crystal Structure Of The Open State Of The Viral Channelrhodopsin Olpvr1
Organism: Organic lake phycodnavirus
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2025-03-12 Classification: MEMBRANE PROTEIN Ligands: RET, NA, 97N, OLC, LFA, GOL |
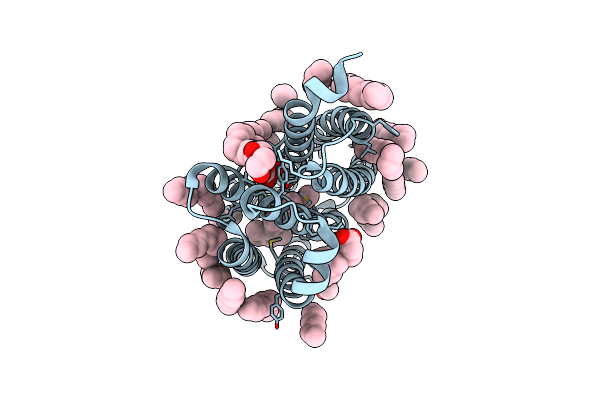 |
Structure Of The Viral Channelrhodopsin Olpvr1 At Low Ph Obtained By Soaking
Organism: Organic lake phycodnavirus
Method: X-RAY DIFFRACTION Resolution:1.41 Å Release Date: 2025-03-12 Classification: MEMBRANE PROTEIN Ligands: RET, 97N, LFA, OLC |
 |
Organism: Subtercola endophyticus
Method: X-RAY DIFFRACTION Resolution:2.46 Å Release Date: 2024-12-11 Classification: MEMBRANE PROTEIN Ligands: LFA, PO4, RET, OLA |
 |
Crystal Structure Of The Cryorhodopsin Cryor2 At Ph 4.6, Type B Crystals, Non-Illuminated State
Organism: Subtercola sp.
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2024-12-11 Classification: MEMBRANE PROTEIN Ligands: LFA, PO4, RET, OLA |
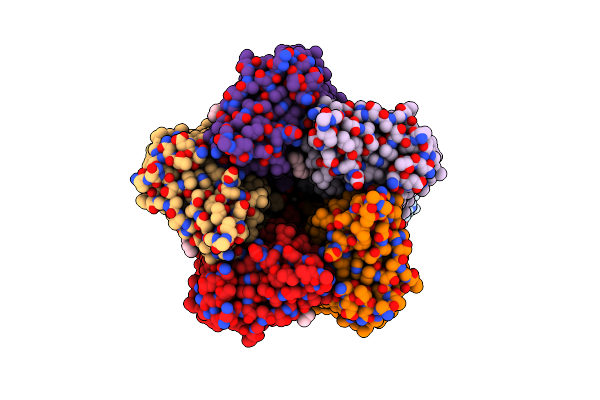 |
Crystal Structure Of The Cryorhodopsin Cryor2 At Ph 4.6, Type B Crystals, Illuminated State
Organism: Subtercola sp.
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2024-12-11 Classification: MEMBRANE PROTEIN Ligands: LFA, PO4, RET, OLA |
 |
Crystal Structure Of The Light-Driven Sodium Pump Ernar In The Monomeric Form At Ph 4.6
Organism: Erythrobacter
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2024-04-24 Classification: MEMBRANE PROTEIN Ligands: LFA, OLA |
 |
Crystal Structure Of The Light-Driven Sodium Pump Ernar In The Monomeric Form At Ph 8.8
Organism: Erythrobacter
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2024-04-24 Classification: MEMBRANE PROTEIN Ligands: LFA, OLA |
 |
Cryo-Em Structure Of The Light-Driven Sodium Pump Ernar In The Pentameric Form At Ph 8.0
Organism: Erythrobacter
Method: ELECTRON MICROSCOPY Resolution:2.63 Å Release Date: 2024-04-24 Classification: MEMBRANE PROTEIN Ligands: LFA, LMT |
 |
Cryo-Em Structure Of The Light-Driven Sodium Pump Ernar In The Pentameric Form At Ph 4.3
Organism: Erythrobacter
Method: ELECTRON MICROSCOPY Resolution:2.50 Å Release Date: 2024-04-24 Classification: MEMBRANE PROTEIN Ligands: LMT, LFA |

