Search Count: 30
 |
Crystal Structure Of Domain-Of-Unknown-Function Duf4867 From Bacillus Megaterium
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-02-26 Classification: ISOMERASE Ligands: FE, NA, CL |
 |
Crystal Structure Of Domain-Of-Unknown-Function Duf4867 From Bacillus Megaterium (Unmodelled Additional Ligand Density At Active Site)
Organism: Priestia megaterium
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-02-26 Classification: ISOMERASE Ligands: FE |
 |
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: EDO, ZN |
 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn From Cupriavidus Pinatubonensis In Complex With Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:2.24 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI |
 |
Crystal Structure Of Hpsn D352A Mutant From Cupriavidus Pinatubonensis In Complex With Nad+
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: NAD, ZN |
 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn From Cupriavidus Pinatubonensis In Complex With Product Analogue (L-Cysteate) And Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI, OCS |
 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn From Cupriavidus Pinatubonensis In Complex With Product (R-Sulfolactate) And Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI, 3SL |
 |
Crystal Structure Of Dhps-3-Dehydrogenase, Hpsn H319A Variant From Cupriavidus Pinatubonensis In Complex With Substrate (R-Dhps) And Nadh
Organism: Cupriavidus pinatubonensis jmp134
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2024-09-25 Classification: OXIDOREDUCTASE Ligands: ZN, NAI, A1AZH, SO4 |
 |
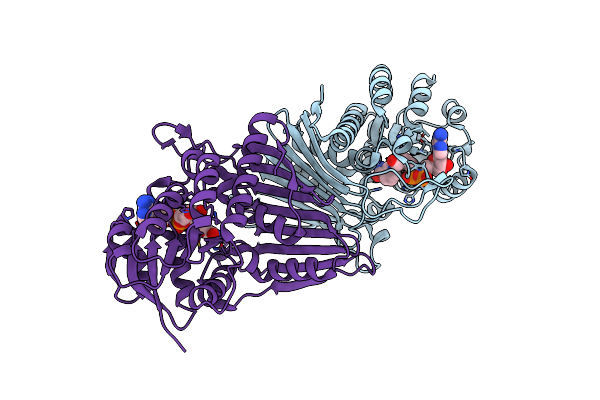
Crystal Structure Of Nad-Dependent Glycoside Hydrolase From Flavobacterium Sp. (Strain K172) In Complex With Co-Factor Nad+ And Sulfoquinovose (Sq)
Organism: Paenarthrobacter ureafaciens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2023-12-27 Classification: HYDROLASE Ligands: NAD, R7R, TRS |
 |
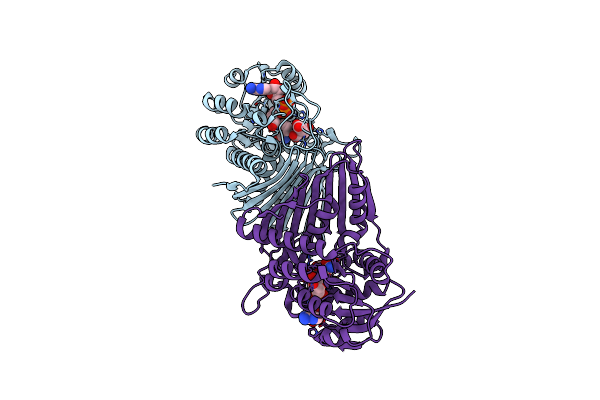
Crystal Structure Of Oxidoreductive Sulfoquinovosidase From Arthrobacter Sp. U41 (Arsqga)In Complex With Co-Factor Nad+
Organism: Arthrobacter sp. u41
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2023-12-27 Classification: HYDROLASE Ligands: NAD |
 |
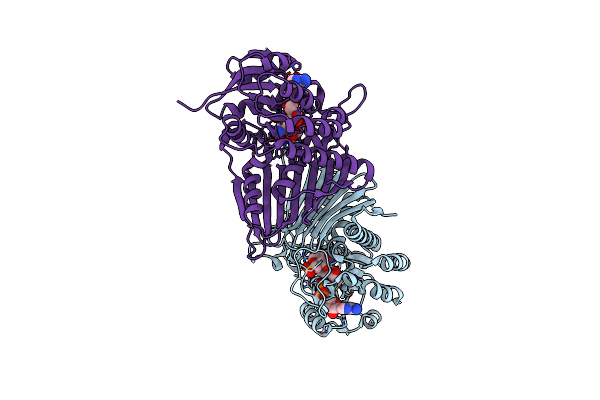
Crystal Structure Of Nad-Dependent Glycoside Hydrolase From Arthrobacter Sp. U41 In Complex With Nad+ Cofactor And Citrate
Organism: Arthrobacter sp. u41
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2023-12-27 Classification: HYDROLASE Ligands: NAD, CIT |
 |
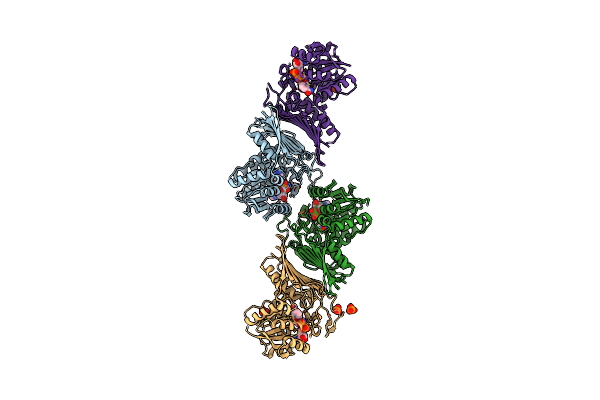
Crystal Structure Of Nad-Dependent Glycoside Hydrolase From Arthrobacter Sp. U41 In Complex With Nad+ And Sulfoquinovose (Sq)
Organism: Arthrobacter sp. u41
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2023-12-27 Classification: HYDROLASE Ligands: NAD, R7R |
 |
Crystal Structure Of Nad-Dependent Glycoside Hydrolase From Flavobacterium Sp. (Strain K172) In Complex With Co-Factor Nad+
Organism: Paenarthrobacter ureafaciens
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2023-12-27 Classification: HYDROLASE Ligands: PO4, NAD |
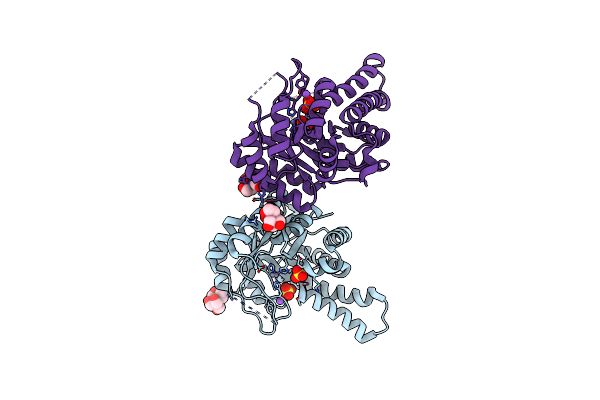 |
Crystal Structure Of Class Ii Sfp Aldolase From Yersinia Aldovae (Yasqia-Zn-So4) With Bound Sulfate Ions
Organism: Yersinia aldovae
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2023-10-25 Classification: LYASE Ligands: SO4, ZN, NA, BTB |
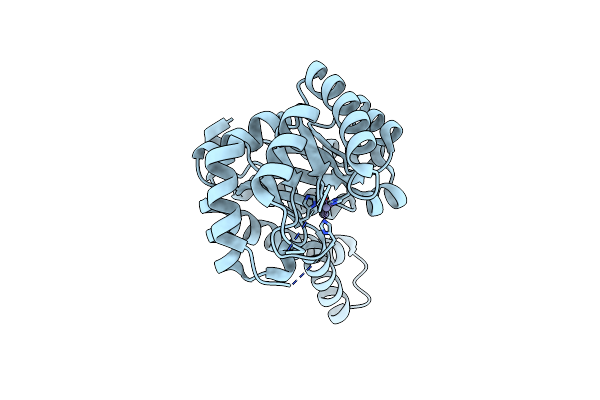 |
Crystal Structure Of Metal-Dependent Classii Sulfofructosephosphate Aldolase (Sfpa) From Hafnia Paralvei Hpsqia-Zn
Organism: Hafnia paralvei
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2023-10-25 Classification: LYASE Ligands: ZN |
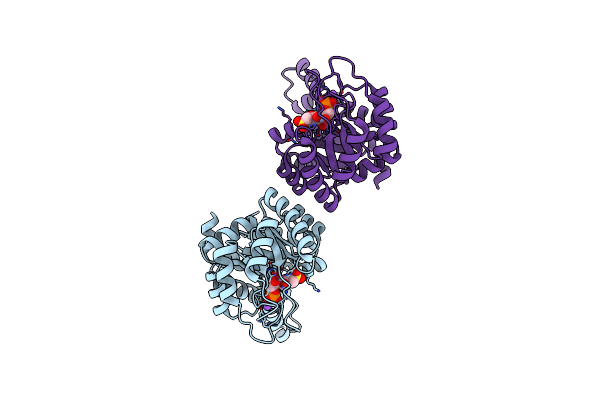 |
Crystal Structure Of Metal-Dependent Class Ii Sulfofructose Phosphate Aldolase From Yersinia Aldovae In Complex With Sulfofructose Phosphate (Yasqia-Zn-Sfp)
Organism: Yersinia aldovae
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2023-10-25 Classification: LYASE |
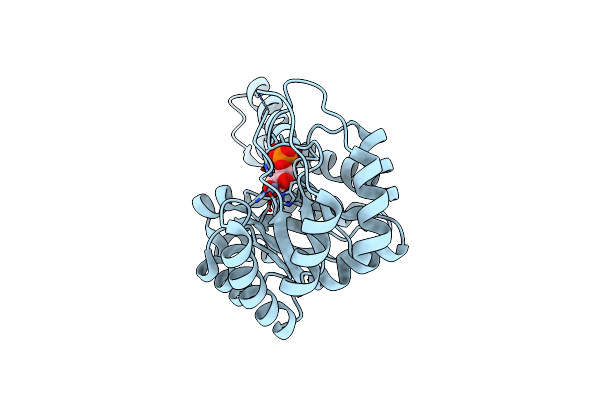 |
Crystal Structure Of Metal-Dependent Class Ii Sulfofructosephosphate Aldolase From Hafnia Paralvei Hpsqia-Zn In Complex With Dihydroxyacetone Phosphate (Dhap)
Organism: Hafnia paralvei
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2023-10-25 Classification: LYASE Ligands: ZN, 13P |
 |
Organism: Amycolatopsis sulphurea
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2023-08-02 Classification: TRANSCRIPTION Ligands: AV8, MG |
 |
Organism: Bacillus subtilis (strain 168)
Method: X-RAY DIFFRACTION Resolution:2.65 Å Release Date: 2018-01-03 Classification: TRANSFERASE Ligands: COA |
 |
Organism: Bacillus subtilis (strain 168)
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2018-01-03 Classification: TRANSFERASE Ligands: COA, GOL, NA |

